Monitoring the rise and fall of the microbiome
Trillions of bacteria live in each person's digestive tract. Scientists believe that some of these bacteria help digest food and stave off harmful infections, but their role in human health is not well understood.
To help shed light on the role of these bacteria, a team of researchers led by MIT associate professor Eric Alm recently tracked fluctuations in the bacterial populations of two research subjects over a full year. The findings, described in the July 25 issue of the journal Genome Biology, suggest that while these populations are fairly stable, they undergo daily fluctuations in response to changes in diet and other factors.
"On any given day, the amount of one species could change manyfold, but after a year, that species would still be at the same median level," says Alm, the Karl Van Tassel Career Development Associate Professor of Biological and Environmental Engineering and senior author of the paper. "To a large extent, the main factor we found that explained a lot of that variance was the diet."
Alm and Lawrence David, an assistant professor at Duke University and the paper's lead author, began the study in 2009, around the same time the National Institutes of Health launched the Human Microbiome Project—an effort to analyze the composition of bacterial communities found in humans, including the gut but also other organs such as the skin, nasal passages, and mouth.
There are a few thousand strains of bacteria that can inhabit the human gut, but only a few hundred of those are found in any given individual, Alm says. For one year, the two subjects in the study collected daily stool samples so bacterial populations could be measured. They also used an iPhone app to track lifestyle factors such as diet, sleep, mood, and exercise, generating a huge amount of data.
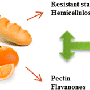
Analysis of this data revealed that dietary changes could produce daily variations in the populations of different strains of bacteria. For example, an increase in fiber correlated with a boost in the populations of Bifidobacteria, Roseburia, and Eubacterium rectale. Four strains—including Faecalibacterium prausnitzii, which has been implicated in protecting against inflammatory bowel disease—were correlated with eating citrus.
During the study, each of the two subjects experienced an event that dramatically altered the gut microbiome. Subject B experienced food poisoning caused by Salmonella, and Subject A traveled to a developing nation, where he experienced diarrheal illness for two weeks.
During Subject B's infection, Salmonella leapt from 10 percent of the gut microbiome to nearly 30 percent. At the same time, populations of bacteria from the phylum Firmicutes, believed to be beneficial to human health, nearly disappeared. After the subject recovered, Firmicutes rebounded to about 40 percent of the total microbiome, but most of the strains were different from those originally present.
"There really wasn't an overall change in the level of Firmicutes. The levels are very different from person to person, but whatever it is about a person that is maintaining a certain level is able to survive a huge insult like that. What's keeping it at that level is a really interesting research question," Alm says, adding that the answer may lie in dietary differences.
Subject A also exhibited severe disruptions to his microbiome during his trip, but once he returned to the United States, it returned to normal. Unlike Subject B's recovery from food poisoning, Subject A's populations returned to their original composition.
The researchers now plan to investigate why microbiome populations tend to return to an average value, specific to the individual host, after going way up or down. They also plan to study how the immune system responds to changes in the microbiome by measuring daily changes in cytokine and hormone levels. To make such follow-up studies easier to do, they are also working on wearable sensors that would collect much of the needed data with less effort required from the subjects.
The ultimate goal, Alm says, is to generate data from individual patients that could be analyzed to produce a personalized monitoring system for people suffering from inflammatory bowel disease or other diseases prone to occasional flare-ups. Such a system could detect when someone is heading for a flare-up and recommend dietary changes that could help avoid it.