Possible new breast cancer gene
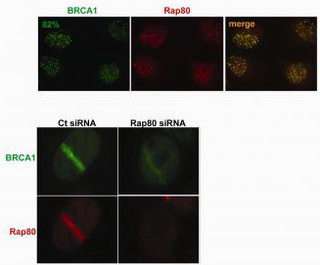
Researchers at the Abramson Family Cancer Research Institute of the University of Pennsylvania and the Dana-Farber Cancer Institute describe in this week’s issue of Science a new candidate breast-cancer susceptibility gene. The Rap80 gene is required for the normal DNA-repair function of the well-known breast cancer gene BRCA1.
Cancer-causing mutations in the BRCA1 protein fail to bind to the Rap80 protein. Consequently BRCA1 is unable to identify DNA damage sites in the genome. When BRCA1 fails to fix DNA damage, cancer-causing mutations accumulate, spawning the development of breast and ovarian malignancies.
"With this current discovery, we have made significant new insights into the molecular mechanism by which BRCA1 recognizes sites of DNA damage that breast-cancer-causing mutated forms of BRCA1 cannot recognize," says co-senior author Roger Greenberg MD, PhD, Assistant Professor of Cancer Biology at Penn. "Now we have gained a partial understanding of the molecular basis between cancer-causing BRCA1 failures to fix DNA damage versus normal BRCA1’s ability to fix DNA damage."
In this study, the researchers found Rap80 binds to the region of the BRCA1 protein that is necessary for recognizing sites of DNA damage. In the 1990s, investigators discovered that BRCA1 was involved in DNA repair by maintaining the normal number and structure of chromosomes. DNA breaks that aren't repaired can lead to cancer by increasing the rate of mutations, cancer-causing changes in the gene sequence.
More specifically, modification of proteins in the cell nucleus – by another protein called ubiquitin – that are tightly bound to DNA are responsible for signaling BRCA1 via Rap80 to action. Rap80 binds to specific types of ubiquitin that concentrate at DNA damage sites, enabling BRCA1 to be recruited to sites of damage.
BRCA1 and BRCA2 mutations account for less than 50 percent of inherited breast cancer. "The genetic basis of breast cancers in other families has been largely unknown," explains Greenberg. "These families aren’t able to make informed choices about screening and treatment, prophylactic or otherwise, the way the BRCA families can."
Researchers from multiple labs, including Penn and those led by co-senior author David M. Livingston at Dana-Farber, are finding that many of these non-BRCA families have mutations in genes that have a relationship with BRCA1. Many of the genes that encode these proteins are also altered in familial breast cancer.
"Thus Rap80, by interacting with a BRCA1 region that is essential for BRCA tumor suppression, now becomes a candidate to investigate as another breast cancer disease gene in families that do not have BRCA1 and BRCA2 mutations, but have a history of breast and/or ovarian cancer," says Greenberg. "In collaboration with other researchers we are currently looking to see if families that have a history of breast cancer, but lack BRCA1 and BRCA2 mutations, have any gene sequence changes in Rap80."
Source: University of Pennsylvania School of Medicine