New clue emerges for cellular damage in Huntington's disease
"Huntington's disease presents an ideal vantage point to study neurodegenerative disease, because we know the misfolded protein that's responsible," says Martin Duennwald, formerly a postdoctoral researcher in the lab of Whitehead Member Susan Lindquist. "But we don't understand how this protein causes cellular damage and death for the neurons that are affected."
In a study published in Genes & Development online on November 17, however, Duennwald and Lindquist report the discovery of a mechanism driven by the misfolded proteins that could be one early trigger for cell death.
In the U.S., about 1 in 20,000 people suffers from Huntington's. Better understanding of the cellular toxicity may allow new therapies for this fatal and incurable disorder.
"This is a diabolical disease, because the misfolded protein interacts with and probably traps many other proteins in the cell," notes Lindquist, who is also a Howard Hughes Medical Institute investigator and a professor of biology at Massachusetts Institute of Technology.
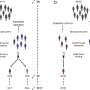
Scientists have long known that Huntington's is driven by a single mutated gene that creates proteins with abnormally long repeats of the amino acid glutamine ("Q"). In certain neurons, these "polyQ-expanded" proteins misfold and clump together, damaging and eventually killing the cells.
But the steps that kick off the process of cell damage and death have remained a mystery, remarks Duennwald, now a principal scientist at Boston Biomedical Research Institute in Watertown, Mass.
In the study, Duennwald first examined what makes polyQ-expanded proteins toxic in yeast. He then performed similar experiments in two kinds of mammalian cells—rat cells that model neurons and mouse striatal cells (from the part of the brain most afflicted in Huntington's).
He found that cells generated with polyQ-expanded fragments quickly showed problems with proteins that had been marked for degradation in the endoplasmic reticulum (ER, a cell component that folds and finalizes proteins). Such proteins were not expelled for tagging and degradation in the cytosol, the intracellular fluid, outside the ER.
"With no garbage disposal, all of a sudden the ER is flooded with proteins that need to be degraded," he says. This breakdown in protein quality control may lead toward cell damage and death.
"We were quite surprised because the ER didn't seem to have any connection with the misfolded proteins in the cytosol," Duennwald adds. "This study tells us to investigate cellular pathways beyond the usual suspects."
He went on to uncover the basis for this breakdown: The polyQ-expanded fragments glom onto the key VCP/Npl4/Ufd1 protein complex that aids in the transport and degradation of the proteins that flunk quality control in the ER. When Duennwald genetically modified cells to over-express two crucial proteins in the protein complex, the toxic effect dropped.
Additionally, his experiments showed that polyQ-expanded proteins avoid a main method by which cells deal with misfolded proteins. Generally, a class of proteins called "chaperone" or "heat shock" proteins move in and either help the misfolded proteins assume their normal shape or help to get rid of them. "Amazingly, polyQ-expanded proteins don't elicit the heat shock response, and that might contribute to their toxicity," Duennwald says.
Such findings may help in eventually treating the disease. The research suggests that activating the cell's protein quality control mechanisms may provide novel and effective strategies for combating Huntington's and other illnesses driven by polyQ-expanded proteins.
Source: Whitehead Institute for Biomedical Research