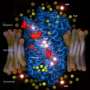
team takes high resolution photo of a K+ channel
Using chemical labeling and mass spectrometry-based techniques, Mark Chance, PhD, director of the Case Western Reserve University School of Medicine Center for Proteomics and Bioinformatics and professor of physiology and biophysics; Sayan Gupta, PhD, instructor at the Case Center for Proteomics and Bioinformatics; and a research team from the University of Oxford, for the first time, were able to take a high resolution picture of the open state of a K+ channel, allowing them to comparatively analyze gating mechanisms important to heart function and nerve signaling - in addition these techniques have already permitted Case Western investigators to gain a deeper understanding of G-protein coupled receptors (GPCRs).
GPCRs, the targets of more than half of all pharmaceutical drugs, are important signaling molecules and help regulate proper channel function. With a new window on the open state of the channel, researchers have an opportunity to better understand the effectiveness of drugs used to treat and prevent cardiovascular diseases.
"The discoveries made by Mark [Chance] and his team have the potential to revolutionize modern medicine as we know it," said Daniel Simon, MD, Chief of Cardiovascular Medicine at University Hospitals Case Medical Center, director of the Case Cardiovascular Center and the Herman K. Hellerstein Professor of Cardiovascular Research at Case Western Reserve School of Medicine. "With an advanced understanding of GPCRs and ion channels comes a better understanding of how modern medicine works, which would theoretically increase pharmaceutical efficiency and enhance the quality of countless cardiovascular-related drugs."
In the study, published in the July edition and featured on the cover of the July issue of Structure, Dr. Chance and his research team discovered the details of the paths by which ions flow through the inwardly-rectifying potassium channel KirBac3.1.The results of this study provide support for a newly proposed gating mechanism of the K+ channel, which if perfected, could offer scientists a new method of probing other important membrane proteins and ion channels that could lead to advancements in the field of cardiovascular medicine. Cardiovascular complications such as stroke and coronary heart disease remain the number one cause of death in all Americans.
To further support his research efforts, Dr. Chance and the Case Western Reserve Center for Proteomics and Bioinformatics received a four-year grant totaling $1.1 million from the National Institute of Biomedical Imaging and Bioengineering (NIBIB). The grant work will be performed in collaboration with Krzysztof Palczewski, PhD, chair of the Department of Pharmacology at Case Western Reserve University, and Dr. Gupta, instructor at the Case Center for Synchrotron Biosciences (CSB) in New York. Dr. Chance is also the director of the CSB, which provides unique facilities to carry out protein footprinting experiments on GPCRs and their complexes.
"Determining the structure of membrane proteins like GPCRs is particularly difficult, while understanding how drugs function to turn them on and off is even harder, but very important to treating major diseases like depression, heart failure, and diabetes," Dr. Chance said of his research. "Our techniques have provided novel insight into addressing these questions and this funding from the NIH will accelerate these studies."
With this funding, Dr. Chance and co-investigators will continue their development of new biotechnology methods to study GPCRs. The aim of their research is as follows:
- To improve their mass spectrometry based structural imaging technology 1,000-fold in an effort to better understand how GCPRs are activated and signal information to protein receptors
- To develop a novel oxygen-18 based water labeling technique to examine the locations and dynamics of structural waters and the exchange properties of bulk water in multiple biological states of interest
- To improve detection efficiency in current protein footprinting experiments in hopes of enhancing the number of amino acids routinely detected
- To develop an algorithmic formula in relation to current footprinting data to determine the outlines of different protein structures
More information: Drs. Chance, Palczewski, Gupta and others have previously published their footprinting/GPCR research in the Proceedings of the National Academy of Sciences.