Researchers hone in on a protein's precise role in disease prevention
Building upon work initiated by University of Alabama scientists, Harvard Medical School researchers, collaborating with UA and others, took another step in cracking the code behind a protein’s precise role in the disease dystonia and, along the way, discovered a new lead for cystic fibrosis research, according to a journal article publishing today.
The research, publishing in this week’s Nature Communications, describes the location and mechanism within a cell where a specific protein, torsinA, normally works to degrade misfolded proteins before those faulty proteins trigger problems associated with the neurological movement disorder called dystonia, said Dr. Guy Caldwell, professor of biological sciences at UA and one of the University’s three co-authors on the article.
“Last year, we started to define that torsinA is functioning within a quality control mechanism, and, in this latest paper, we hone in on the precise portion of cellular quality control that it seems to be managing,” Caldwell said.
The new findings suggest that in one area of cells, the endoplasmic reticulum – which Caldwell likens to a cell’s post office, a place where the cell’s proteins are packaged before transport to other parts of the cell – torsinA plays a key role.
In the absence of torsinA’s quality control function, the misfolding of proteins within the endoplasmic reticulum continues, leading to cell stress from protein aggregation and, eventually, to neuron malfunction, the researchers said. This neuron malfunction leads to dystonia.
In another aspect of the study, the researchers show a potentially promising relationship between torsinA activity and cystic fibrosis, a life-threatening lung disease said to be the most common disease among young Caucasians. Scientists previously knew, Caldwell said, that cystic fibrosis results from the deletion of a single amino acid within the protein called cystic fibrosis transmembrane conductance regulator, or CFTR, that must be properly folded within the endoplasmic reticulum.
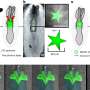
The new study shows that, in both the C. elegans nematode roundworms that the Caldwell Lab uses as animal models, and in human cells, that torsinA can selectively degrade the misfolded CFTR protein, while a mutant version of torsinA is less able to do so.
“If you are thinking about cures for cystic fibrosis, what you really want is to be able to manage the amount of CFTR protein being sent through the endoplasmic reticulum. Regulating torsinA activity has the potential to modulate that,” Caldwell said.
The Caldwell Lab was recently awarded a one-year, renewable grant from the National Institutes of Health, via the UAB Cystic Fibrosis Research Center, to continue its cystic fibrosis research.
Likewise, Caldwell said research into dystonia has proceeded rapidly over the last decade and the latest step could prove vital in the development of therapeutic strategies.
“Until you can understand how a protein works in the cell, you can’t really develop biological assays, or tests, for the way in which it can be modified, let alone find drugs that can modify it,” he said.
Researchers from UA’s Caldwell Lab were the lead authors of a 2010 paper, also in collaboration with Harvard, publishing in Human Molecular Genetics about the role of torsinA, as well as a 2003 cover article in the same journal where torsinA’s therapeutic potential was initially linked to disorders involving management of protein misfolding.
About 500,000 people in North American have dystonia, a disorder that can cause repetitive, painful spasms and can affect one or many parts of the body. Patients with early-onset generalized dystonia have one good copy of the gene DYT1, and one problematic copy, in their DNA. It’s this gene that contains the information to make torsinA.
The multi-institution effort published in Nature Communications was led by Dr. Xandra Breakefield, a professor in the department of neurology at Harvard Medical School and a geneticist at Massachusetts General Hospital. The research was done in both animal models and via skin cells obtained directly from dystonia patients.