For those with the rarest diseases, genomes can yield answers
For many of us, having our genomes in hand today isn't likely to make any profound difference in our lives, at least not when it comes to our health. But for children and their families affected by rare and mysterious genetic diseases, early indications are that it's a completely different story, thanks to the efforts of two teams of geneticists at Duke Medicine.
When Nico Katsanis arrived at Duke three years ago, he knew he wanted to do something transformative, by finding ways to bridge basic science and clinical medicine. That idea began to take shape in early 2011 in the form of the Duke Task Force for Neonatal Genomics. The plan: to bring patients, researchers and clinicians together to apply the latest genomic technologies to the diagnosis of newborns and very young children with some of the most challenging clinical disorders.
"We really have an opportunity to bring together everything that we've learned – in molecular biology, cell biology, biochemistry – and put that together with cutting edge genomics and actually try to make a difference in real-time in people's lives," Katsanis said.
It was an effort that began as slowly as it did quietly. In the first six months, one family signed up. The next month, it was two or three. A year later, the task force has sequenced the full exomes (all of the protein-coding regions in the genome) of 15 children and is enrolling new families weekly.
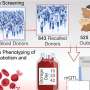
In the beginning, even getting a phlebotomist at the right place and the right time was hard. Now, they've built the right team and an efficient process. When a Duke clinician thinks that they may have a case, they send word by email to the group. Decisions about entry in the study are made by consensus; approval is needed from each arm of the task force: the clinicians, the basic scientists and the genome ethics and policy experts. If it's a go, the referring physician introduces the family to a genetic counselor. Once the family has consented, DNA samples are sent off to research partners at Baylor College of Medicine.
Gone fishing
So far, the DNA evidence has led to a diagnosis with reasonably high confidence based on the scientific literature about 40 percent of the time – a satisfying result, especially considering that these are most often newborns; they may not yet display the full spectrum of symptoms and they haven't undergone years of testing in the diagnostic odyssey that many families endure.
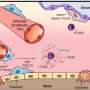
For the other 60 percent, the sequence is just the start. The task force might find a handful of genes that look suspicious for one reason or another, but no obvious diagnosis. That's when the real work begins. Their goal is to functionally investigate each and every one of those mutations. Often that means that Assistant Professor of Pediatrics Erica Davis and the rest of the team at the Center for Human Disease Modeling that Katsanis directs will test those mutant human genes in zebrafish embryos one by one or in combination, tos ee how they affect development of the fish and its organs and how that compares to the child's condition. Other times, zebrafish aren't the answer.
"That's when we have to get creative," Davis says. They are enlisting collaborators from Duke and at other institutions with the appropriate expertise to test things like, for example, the effects of a mutant channel on cell membrane electrophysiology. In one case, they've even gone to the level of inducing stem cell-like cells.
"We don't start with any preconceived ideas," Katsanis said. "We find all of the genes that are consistent with causality and we test them. It's tedious and expensive and time consuming."
So far, it also works. Davis reported in a plenary presentation to the American Society of Human Genetics that they have a strong candidate for diagnosis in about 90 percent of cases. Michael Cotten, a pediatric neonatologist, says those answers are likely to influence care.
Sometimes the evidence may lead to a telling clinical test that wouldn't have been ordered otherwise, other times to a different therapeutic strategy. If the messages that are already filling email inboxes are any indication, it's making a real difference for the families, who are treated as equal partners along the way.
Sequencing clinic
Another Duke effort is offering answers to children and families who haven't been so lucky and, after years of conventional clinical and medical genetic testing, still don't have their answers to challenging medical conditions. David Goldstein, director of the Center for Human Genome Variation, and Vandana Shashi, who leads the Genome Sequencing Clinic, meet several times a year to evaluate children with complex medical cases involving unexplained developmental delay, intellectual disability or birth defects and to sequence their genomes in Goldstein's laboratory. Earlier this year, Goldstein and Shashi reported in the Journal of Medical Genetics that they had found a diagnosis for seven of the first twelve children with genomes in hand.
"These are the hardest of the hard cases and we find we can resolve them about half the time," Goldstein said. "Complete genomes provide that answer. It's the clearest example I know of how important genomics can be in rare diseases."
These clinical efforts are buoyed by a growing number of human exomes being sequenced in a research mode in the IGSP's Genome Sequencing and Analysis Core Resource. According to Greg Wray, director of the resource, dozens of samples are at some stage of sequencing for investigators in the Departments of Medicine, Anesthesiology and Biomedical Engineering, with more on the way.
Wray said that their new exome sequencing service is available to all and represents a partnership between the IGSP's DNA Microarray Core Facility, which does the initial exome capture, and the Genome Sequencing Core. "Our vision is that the whole can be greater than the sum of its parts," he said.
Certain progress amidst uncertainty
Genome sequences can also reveal whether pathogenic mutations were passed from parent to child or are brand new and unlikely to recur in a subsequent pregnancy. That information is critical when it comes to genetic counseling for families, although there are challenges in communicating uncertainty, Goldstein says.
Those are issues that the neonatal task force, too, has largely yet to face as the families get through their initial medical crises and consider what else their genome information might mean. That's where IGSP policy experts Misha Angrist and Sara Katsanis come in with efforts to explore those very personal issues.
For pediatric urologist John Weiner, the task force embodies "why I came back to Duke. I wanted to work with basic scientists and to meld these two worlds." Obstetrician Amy Murtha is already looking ahead to a future in which doctors might have these answers in hand even before a new baby with serious health problems is born.
"My job is trying to push the envelope back into pregnancy," she says. "We aren't there yet, but that's the direction we need to go. I think Duke is in a position to do that well and to help establish the guidelines."