Hidden strains of HPV found in 'virus-negative' genital warts
There are 170 established HPV types. Cancerous human papillomavirus (HPV) viruses are the main cause of cervical cancer, and are found in close to 100% of cervical tumors.
Cervical cancer and genital warts are caused by HPV. However, testing for the virus using standard techniques can sometimes give a negative result—in these cases, the condylomas are called 'virus-negative' warts.
In a new study published in Virology, researchers assessed the DNA found in samples taken from 40 patients with 'virus-negative' genital warts. Through a general DNA sequencing approach, the researchers showed that several of the negative samples did in fact contain HPV DNA.
This means that virus-negative warts can harbor small amounts of more distantly related viruses that escaped previous detection. According to the research, there is a diverse pool of previously unknown HPV types that infect humans and are detectable on genital warts.
The findings have implications for the knowledge of diversity of HPV types, as these viruses are currently undetectable using traditional testing methods.
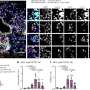
Ten pools of four samples taken from virus-negative warts were tested using genetic material straight from the patient—including viral, bacterial and human DNA. Five of the pools contained HPV, and three of these contained new strains of the virus.
Altogether, 1337 pieces of HPV-related DNA were detected, representing 23 new types of HPV, 10 established types of HPV and two known HPV DNA sequences.
This new style of testing has highlighted previously unknown forms of the virus. As such, we learn more about the evolution of different HPV types. It is possible that the previously unknown forms of the virus do not cause condyloma but may be secondary invaders of condyloma.
More information: This article is "Metagenomic sequencing of "HPV-negative" condylomas detects novel putative HPV types" by Hanna Johansson, Davit Bzhalava, Johanna Ekström, Emilie Hultin, Joakim Dillner, and Ola Forslund (DOI: 10.1016/j.virol.2013.01.023) and appears in Virology.