Mapping the DNA sequence of Ashkenazi Jews
Led by Itsik Pe'er, associate professor of computer science at Columbia Engineering, a team of researchers has created a data resource that will improve genomic research in the Ashkenazi Jewish population and lead to more effective personalized medicine. The team, which includes experts from 11 labs in the New York City area and Israel, focused on the Ashkenazi Jewish population because of its demographic history of genetic isolation and the resulting abundance of population-specific mutations and high prevalence of rare genetic disorders. The Ashkenazi Jewish population has played an important role in human genetics, with notable successes in gene mapping as well as prenatal and cancer screening. The study was published online on Nature Communications today.
"Our study is the first full DNA sequence dataset available for Ashkenazi Jewish genomes," says Pe'er, who is also a co-chair of the Health Analytics Center at Columbia's Institute for Data Sciences and Engineering, as well as a member of its Foundations of Data Science Center. "With this comprehensive catalog of mutations present in the Ashkenazi Jewish population, we will be able to more effectively map disease genes onto the genome and thus gain a better understanding of common disorders. We see this study serving as a vehicle for personalized medicine and a model for researchers working with other populations."
To help in his hunt for disease genes, Pe'er founded The Ashkenazi Genome Consortium (TAGC) in September 2011 with Todd Lencz, an investigator at The Feinstein Institute for Medical Research, director of the Laboratory of Analytic Genomics at the Zucker Hillside Hospital, and associate professor of molecular medicine and psychiatry at the Hofstra North Shore-LIJ School of Medicine. The other TAGC members, who are providing expertise in the diseases they are studying, are:
- Gil Atzmon, associate professor of medicine and genetics, Albert Einstein College of Medicine (genetics of longevity and diabetes);
- Lorraine Clark, associate professor of clinical pathology and cell biology and co-director, Personalized Genomic Medicine Laboratory, Columbia University Medical Center, Laurie Ozelius, associate professor at Icahn School of Medicine at Mount Sinai, and Susan Bressman, chair of neurology at Mount Sinai Beth Israel (Parkinson's disease and related neurological phenotypes);
- Harry Ostrer, professor of pathology, genetics, and pediatrics, Albert Einstein College of Medicine (radiogenomics, cancers and rare genetic disorders);
- Ken Offit, chief of clinical genetics at Memorial Sloan Kettering Cancer Center (breast, ovarian, colon and prostate cancers, lymphoma);
- Inga Peter, associate professor of genetics and genomic sciences, and Judy Cho, professor of medicine and professor of genetics and genomic sciences, both at The Mount Sinai Hospital (inflammatory bowel disease); and
- Ariel Darvasi, vice-dean of The Faculty of Life Sciences at The Hebrew University of Jerusalem (multiple diseases).
Before the TAGC study, data was available for a limited number of DNA markers (only approximately one in every 3000 letters of DNA) that are mostly common in Europeans. The TAGC researchers performed high-depth sequencing of 128 complete genomes of Ashkenazi Jewish healthy individuals. They compared their data to European samples, and found that Ashkenazi Jewish genomes had significantly more mutations that had not yet been mapped. Pe'er and his team analyzed the raw data and created a comprehensive catalog of mutations present in the Ashkenazi Jewish population.
The TAGC database is already proving useful for clinical genomics, identifying specific new mutations for carrier screening. Lencz explains: "TAGC advances the goal of bringing personal genomics to the clinic, as it tells the physician whether a mutation in a patient's genome is shared by healthy individuals, and can alleviate concerns that it is causing disease. Without our work, a patient's genome sequence is much harder to interpret, and more prone to create false alarms. We have eliminated two thirds of these false alarms."
The TAGC study further enables more effective discovery of disease-causing mutations, since some genetic factors are observable in Ashkenazi individuals but essentially absent elsewhere. Moreover, the demography of the Ashkenazi population, the largest isolated population in the U.S., enables large-scale recruitment of study patients and hence more genetic discoveries than in other well-known isolated populations like the Amish and Hutterites locally, or the Icelanders overseas. The researchers expect that medical insights from studies of specific populations will also be relevant to general populations as well.
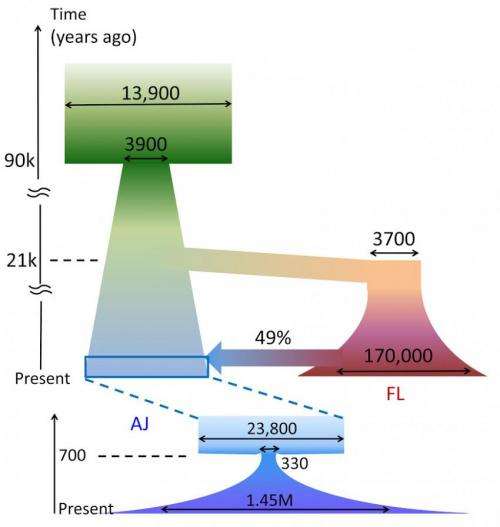
The TAGC team's findings also shed light on the long-debated origin of Ashkenazi Jews and Europeans. The genetic data indicates that the Ashkenazi Jewish population was founded in the late medieval times by a small number, effectively only hundreds of individuals, whose descendants expanded rapidly while remaining mostly isolated genetically.
"Our analysis shows that Ashkenazi Jewish medieval founders were ethnically admixed, with origins in Europe and in the Middle East, roughly in equal parts," says Shai Carmi, a post-doctoral scientist who works with Pe'er and who conducted the analysis. "TAGC data are more comprehensive than what was previously available, and we believe the data settle the dispute regarding European and Middle Eastern ancestry in Ashkenazi Jews. In addition to illuminating medieval Jewish history, our results further pave the way to better understanding European origins, millennia before. For example, our data provides evidence for today's European population being genetically descendant primarily from late mid-eastern migrations that took place after the last ice age, rather than from the first humans to arrive to the continent, more than 40,000 years ago."
The researchers note that their results also raise a number of hypotheses regarding Ashkenazi origin, which they have now begun exploring. They are looking at, for example, the identity of the Ashkenazi founders: who are the modern populations most similar to them in Europe and in the Middle East? When did the admixture process take place and to what extent was that process sex-biased, where men with one origin mated with women from another?
Pe'er adds, "Equally important to us is making our data available to the entire research community. We've released it to public-access databases and fully expect the creativity of the scientific world to come up with additional uses for the data. What's especially gratifying is the idea that our work will pave the way for personalized genomics in other populations as well."
Pe'er and his TAGC colleagues are already working on their next step: to study specific diseases in the Ashkenazi population, including schizophrenia, Parkinson's, Crohn's, diabetes, and cancer, as well as other inherited traits such as longevity. They are also working with additional collaborators, including the New York Genome Center, to sequence approximately 500 more Ashkenazi genomes to make the catalog of mutations even more comprehensive.
More information: Sequencing an Ashkenazi reference panel supports population-targeted personal genomics and illuminates Jewish and European origins, Nature Communications 5, Article number: 4835 DOI: 10.1038/ncomms5835