Sorting bloodborne cancer cells to better predict spread of disease
For most cancer patients, primary tumours are often not the most deadly. Instead, it is the metastatic tumours - tumours that spread from their original location to other parts of the body - that are the cause of most cancer deaths.
The catalysts behind the formation of these deadly metastatic tumours are believed to be cancer cells that are launched into the bloodstream from the original site of the cancer. Researchers are very interested in leveraging these circulating tumour cells, or CTCs, which have the potential to allow the properties of a tumour to be better understood without a biopsy, and may also help physicians recognize how aggressive a tumour is and whether it is likely to cause metastatic disease.
However, not all CTCs in a given patient are alike. Recent discoveries have shown that CTCs are highly heterogeneous - with individual cancer cells possessing very different molecular characteristics - and that only a small subset of these cells actually possess the metastatic potential to spread the disease throughout the body.
Current technologies exist that allow these circulating cells to be captured from the blood of cancer patients, but they are not well equipped to differentiate between the various CTCs present in the blood sample. Instead, they simply count the number of CTCs in a patient sample, rather than identifying the cells that possess the highest metastatic potential. As a result, these tools are less than ideal as they are only able to provide general information on the levels of CTCs rather than a more focused understanding of the disease and its aggressiveness.
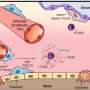
Researchers at the Leslie Dan Faculty of Pharmacy at the University of Toronto have developed a new device that provides a way to visualize the heterogeneity of CTCs, and have published their findings in the leading Chemistry journal Angewandte Chemie. Using nanoparticles to tag cells, this device sorts the CTCs collected in a sample into discrete subpopulations based on the phenotype of the cells, and provides a snapshot of the nature of the tumour cells present in patients' blood.
"Recognizing that characterizing the phenotype of circulating tumour cells is more useful for cancer management than quantitating the cells present in a blood sample, we set out to devise a method that would allow us to capture and distinguish between these cells," notes Professor Shana Kelley of the Leslie Dan Faculty of Pharmacy. "In the lab, we were able to demonstrate that the tool was not only highly effective at differentiating these cells, but also proved to be more sensitive than the current leading methods of cellular sorting."
Partnering with collaborators at the Sunnybrook Health Sciences Centre and the London Health Sciences Centre, the researchers collected samples from prostate cancer patients to test the efficacy and ability of the diagnostic platform.
"Through this study, over 20 patients with localized prostate cancer were tested," notes Dr. Robert Nam, Ajmera Chair in Urologic Oncology and Head of Genitourinary Oncology, at Sunnybrook's Odette Cancer Centre . "Interestingly, significant levels of these circulating tumour cells were observed in all of the patients. Even more intriguing was the observation of very different subpopulation profiles across this group of patients that all received very similar clinical diagnoses, indicating that molecular-‐level difference may exist in the patients' tumours."
While this study only involved a small number of patients, further validation is planned with several other cancers, including breast, colon, ovarian, lung, and pancreatic cancer.
"Ultimately, we believe that this sensitive technology possesses the potential to provide more useful information about these cells, leading to better diagnoses and improved patient outcomes," notes Dr. Kelley.
"As a result, we are excited to pursue new research opportunities in an effort to more accurately and less invasively diagnose and improve the health outcomes for cancer patients."