Researchers map 'switches' that shaped the evolution of the human brain
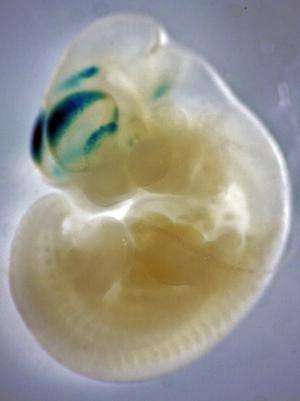
Thousands of genetic "dimmer" switches, regions of DNA known as regulatory elements, were turned up high during human evolution in the developing cerebral cortex, according to new research from the Yale School of Medicine.
Unlike in rhesus monkeys and mice, these switches show increased activity in humans, where they may drive the expression of genes in the cerebral cortex, the region of the brain that is involved in conscious thought and language. This difference may explain why the structure and function of that part of the brain is so unique in humans compared to other mammals.
The research, led by James P. Noonan, Steven K. Reilly, and Jun Yin, is published March 6 in the journal Science.
In addition to creating a rich and detailed catalogue of human-specific changes in gene regulation, Noonan and his colleagues pinpointed several biological processes potentially guided by these regulatory elements that are crucial to human brain development.
"Building a more complex cortex likely involves several things: making more cells, modifying the functions of cortical areas, and changing the connections neurons make with each other. And the regulatory changes we found in humans are associated with those processes," said Noonan, associate professor of genetics, an investigator with the Kavli Institute for Neuroscience, and senior author of the study. "This likely involves evolutionary modifications to cellular proliferation, cortical patterning, and other developmental processes that are generally well conserved across many species."
Scientists have become adept at comparing the genomes of different species to identify the DNA sequence changes that underlie those differences. But many human genes are very similar to those of other primates, which suggests that changes in the way genes are regulated—in addition to changes in the genes themselves—is what sets human biology apart.
Up to this point, however, it has been very challenging to measure those changes and figure out their impact, especially in the developing brain. The Yale researchers took advantage of new experimental and computational tools to identify active regulatory elements—those DNA sequences that switch genes on or off at specific times and in specific cell types—directly in the human cortex and to study their biological effects.
First, Noonan and his colleagues mapped active regulatory elements in the human genome during the first 12 weeks of cortical development by searching for specific biochemical, or "epigenetic" modifications. They did the same in the developing brains of rhesus monkeys and mice, then compared the three maps to identify those elements that showed greater activity in the developing human brain. They found several thousand regulatory elements that showed increased activity in human.
Next, they wanted to know the biological impact of those regulatory changes. The team turned to BrainSpan, a freely available digital atlas of gene expression in the brain throughout the human lifespan. (BrainSpan was led by Kavli Institute member Nenad Sestan at Yale, with contributions from Noonan and Pasko Rakic, a co-author on this study.) They used those data to identify groups of genes that showed coordinated expression in the cerebral cortex. They then overlaid the regulatory changes they had found with these groups of genes and identified several biological processes associated with a surprisingly high number of regulatory changes in humans.
"While we often think of the human brain as a highly innovative structure, it's been surprising that so many of these regulatory elements seem to play a role in ancient processes important for building the cortex in all mammals, said first author Steven Reilly. "However, this is often a hallmark of evolution, tinkering with the tools available to produce new features and functions."
Next, Noonan and colleagues plan to investigate the function of some of the regulatory changes they identified by introducing them into the mouse genome and studying their effects on mouse brain development.
More information: Evolutionary changes in promoter and enhancer activity during human corticogenesis, Science, www.sciencemag.org/lookup/doi/ … 1126/science.1260943