Study validates effectiveness of genomic test for lung cancer detection
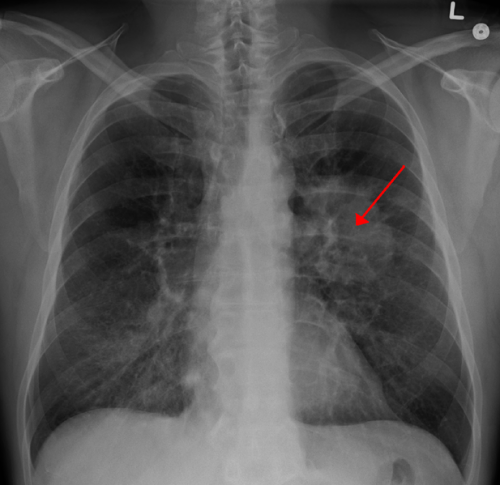
A new test co-developed by a Boston University School of Medicine (BUSM) researcher will allow patients suspected of having lung cancer to be subjected to fewer and less-invasive tests to determine if they have the disease.
"We are seeing an increase in the number of lesions suspicious for lung cancer found on chest imaging of current and former smokers. In the past, these patients have been subjected to invasive tests when traditional bronchoscopy tests prove inconclusive. Today's announcement provides physicians and patients with an additional piece of scientifically reliable information to consider when determining their next diagnostic step," said senior author Avi Spira, MD, MSc, professor of medicine, pathology and bioinformatics at BUSM.
Researchers have found that a genomic biomarker can accurately determine the likelihood of a lung lesion being malignant. The findings that appear online in the New England Journal of Medicine are from two large, prospective, multicenter studies called Airway Epithelium Gene Expression in the Diagnosis of Lung Cancer (AEGIS) I and II. These findings will allow physicians to confidently identify patients who are at low probability for having lung cancer thus sparing them from costly and risky procedures.
The Impact
"While the test itself is simple, the science behind it is remarkable," added Spira who also is the Alexander Graham Bell Professor in Health Care Entrepreneurship at BUSM. Previous work by Spira found that the pattern of gene activity in cells lining the upper respiratory tract can identify cancer that is developing deeper in the lung. "The ability to test for molecular changes in this 'field of injury' allows us to rule out the disease earlier without invasive procedures. Conceptually, this may have significant implications for other diseases."
Study Details
The study involved 639 patients (298 in AEGIS I and 341 in AEGIS II) at 28 sites in the United States, Canada and Ireland who were undergoing bronchoscopy, a common nonsurgical procedure to assess lung lesions for cancer. Using airways cells collected by the bronchoscopy, the researchers found this genomic test, when evaluated with the bronchoscopy, had a combined sensitivity of 97 percent for detecting lung cancer, compared to 75 percent for bronchoscopy alone.
"This study validates the effectiveness of the bronchial genomic biomarker among those undergoing bronchoscopy in two independent groups. We found that it has high sensitivity across different sizes, locations, stages and cell types of lung cancer," added Spira. "The combination of the biomarker and bronchoscopy has a sensitivity of 96 percent and 98 percent in the AEGIS-1 and AEGIS-2 groups, respectively."
An estimated 250,000 patients undergo a bronchoscopy for suspected lung cancer each year with approximately 40 percent producing non-diagnostic results. This can lead to invasive procedures such as transthoracic needle biopsy or surgical lung biopsy that are risky and expensive. "In intermediate risk patients with a non-diagnostic bronchoscopy, a negative genomic test warrants consideration of a more conservative diagnostic approach that could reduce unnecessary invasive testing in patients without lung cancer. We hope to improve the diagnostic work up for lung cancer by reducing patient anxiety, performing fewer unnecessary procedures and ultimately saving valuable healthcare resources and money," Spira said.