Mining critical care data to support real-time clinical decision

In the intensive care units found in leading hospitals, just about every aspect of human physiology is continually monitored and reported. Critical care clinicians synthesize all these inputs in real-time, but given the high volume of data, some important signals may get lost in the mix. Within a few days, most of the data are deleted altogether, preventing further study.
"Physicians coming into intensive care have to screen a lot of data and synthesize it in their heads," says Thomas Heldt, the Helmholtz Career Development Professor with MIT's Institute for Medical Engineering and Science and an assistant professor of electrical and biomedical engineering. "Some of this data comes at a very high temporal resolution, while data concerning aspects like medications and lab results comes much less so. In our work, we try to relate the data high-resolution streams to mathematical models of the underlying physiology to identify model parameters that have clear physiological significance but may not be easily measureable directly. Our goal is to provide the physician with a more complete picture."
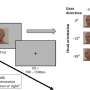
Heldt's principal interest lies in physiological waveforms, such as electrocardiograms or arterial blood pressure measurements. In a typical ICU, there can be about 10 simultaneous streams recorded, digitized at 100 samples per second or more, from the heart, brain, and vascular and respiratory systems. "Some of this data contains redundant information, but this redundancy can be helpful, letting you compare and see when information is diverging," says Heldt. "Having this data available allows us to test out our mathematical modeling approaches with a lot of data."
Heldt is initially focusing on developing modeling techniques that can tease out new signals from conventional measurements, thereby potentially replacing procedures that are costly or dangerous. "If we understand the underlying physiology, we might be able to estimate parameters that cannot be easily measured," says Heldt. "By leveraging an understanding of the underlying physiology, we hope to estimate parameters noninvasively and continuously in a patient specific manner."
Some of the high-risk measurements are found in neurocritical care. "Because the brain is encapsulated by the skull, it is difficult to get at," says Heldt, who in 2014 established the Integrative Neuromonitoring and Critical Care Informatics Group at MIT. "Aside from measuring electrical signals, real-time monitoring of the brain is fairly limited."
In order to measure intracranial pressure (ICP) in patients with severe traumatic brain injury, for example, neurosurgeons drill a hole into the skull and advance a catheter toward the center of the brain. "It's a very invasive procedure that you would only attempt in severely sick patients," says Heldt.
As an alternative, Heldt's group is attempting to estimate ICP by synthesizing data from two easily obtainable waveform measurements: arterial blood pressure and transcranial Doppler based measurements of arterial blood flow velocity. "Our model is relating the physiology of intracranial pressure to these measurements to help us estimate the pressure," Heldt says.
The core challenge here is the mathematical problem of "building a model at the right size that allows us to robustly estimate the parameters," Heldt says. "In some of our work, we use machine learning and data analytic tools to understand the relations between different data streams and between those streams and the patient outcome, but for the most part we try to incorporate our understanding of the underlying physiology into the analysis tools. We can represent the physiology mathematically through differential equation or algebraic equations, and then try to interpret the data within the context of those mathematical relationships. My goal is to try to incorporate as much of that knowledge into the decision making process, modeling, and analysis, as possible."
One challenge in synthesizing medical data from multiple inputs is the vast difference in time scales even within a single type of measurement. For example, while electrocardiograms must be analyzed at scales of hundreds of samples per second, physicians also need to analyze trends over time, sampled in hours or even days. "Our models break down the time scales into small chunks," Heldt says. "The parameters we estimate vary from one time window to the next, so we can see a trend in their evolution."
Perhaps the most difficult task in the neurocritical project is not mathematical, but technological: capturing the data for analysis. "Intensive care units are not set up to archive data," says Heldt. "The waveforms you see on the monitors are only kept for about 72 hours."
To identify new archival procedures, Heldt is working with researchers at Boston's Children Hospital, Beth Israel Deaconess Medical Center, and Boston Medical Center—he has courtesy appointments at all three hospitals. Developing archival systems is difficult because "no two intensive care units in Boston are the same," Heldt says. "Ideally, you validate against different patient conditions, pathologies, age ranges, and hospitals. You also have to demonstrate that the measurements are no worse than the invasive measurements."
Device miniaturization and the road to wearables
Heldt's ambitions for his clinical data research do not stop at the ICU, or even the hospital. "We started with intracranial pressure because that's where the invasive measurement is available for validation," he says. "Once our approach is validated, we hope to migrate out of these highly specialized cases to a much bigger patient pool."
To expand noninvasive intracranial pressure (ICP) monitoring to the larger patient population, clinicians still need arterial pressure and blood flow measurements, but not all the other data collected in critical care. Heldt is counting on new wearable, or at least smaller and more affordable, medical sensor devices to help expand ICP monitoring to the outpatient clinic or possibly even the home. Heldt's group is collaborating with MIT professors Charles Sodini and Harry Lee, as well as MIT's Medical Electronic Device Realization Center.
"Typical ultrasound equipment to estimate the velocity of blood flow in the brain can be the size of a small suitcase," Heldt says. "It requires the patient to wear a big head frame, and you need an operator to steer the ultrasound beam to the right target location. By miniaturizing the electronics and improving the algorithms, there's no need for an operator. We should make it as easy as putting on an ECG electrode so you could use this in a doctor's office, or even at home. The skill of the operator would be encoded in the electronics and associated algorithms."
"We think we can make these devices operator independent, cheaper, and therefore much more widely available and useful," Heldt continues, adding, "I see a lot of opportunity to reinvent medical devices for less expensive care."
Heldt is also looking to improve his models by integrating a growing body of clinical data. "We've drawn insights from clinical databases in trying to understand relationships among data and between the data and particular conditions," says Heldt. "We've been involved in developing big ICU databases, such as the Multiparameter and Intelligent Monitoring in Intensive Care database developed by my MIT PhD advisor Roger Mark."
Eventually, the models might advance to the level where they could provide clinical decision support. "Clinicians are very good at discarding information they think is not relevant and holding onto information that really is important," says Heldt. "That has traditionally been very difficult to teach a machine to do, so we're still exploring to what extent computers can help with all these decision processes."
Bringing modeling to neonatal care
In a more recent project, Heldt is collaborating with clinicians at Beth Israel Deaconess on the study of clinical data from neonates, in particular extreme low-birth-weight preterm neonates. "We are looking at where their oxygen levels should be targeted vs. how much they actually get," says Heldt. "That can feed back into the clinical care process, so a performance metric can be developed."
Heldt's research has benefited greatly from the Boston medical community. "We are happy to be close to the leading hospitals in the world," he says. "The patient population, volume, and collaborative research culture at these hospitals is very conducive to this kind of research. Some of our best work has been initiated by clinicians who came to us with their problems. Those are the best kinds of collaborations. You combine an expert in the field who is very invested in how the patients are treated, with people like us with the analytical expertise to examine the data creatively and to build new devices."
More information: Multiparameter and Intelligent Monitoring in Intensive Care database: mimic.physionet.org/