Bacteria's own genome becomes food safety tool
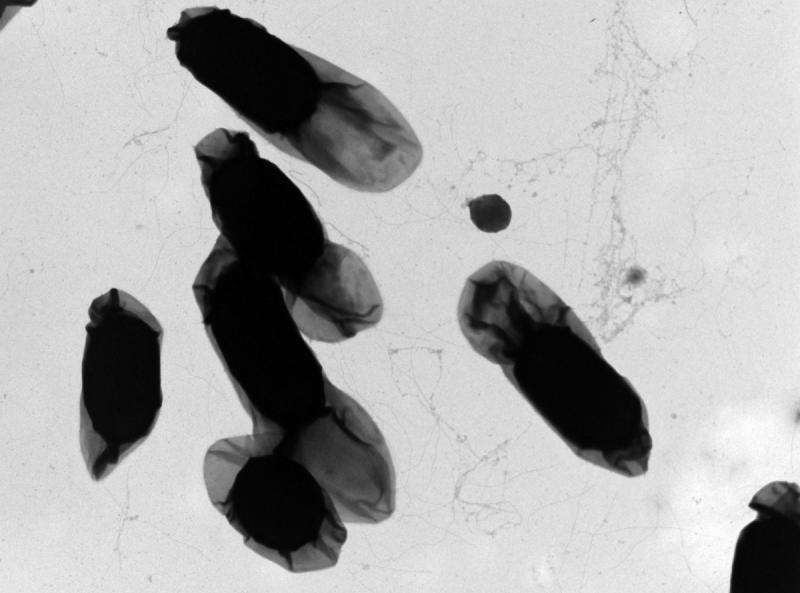
Bacillus cereus – a common food bacterium – can no longer hide. The food industry has a new tool for identifying specific isolates behind foodborne illness that utilizes the bacteria's own genomes, reports Cornell food scientists in the journal BMC Genomics, Aug. 8.
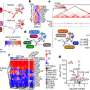
"Examining the whole genome of the B. cereus group is a more reliable tool for identifying risks associated with the presence of these bacteria in our food," said lead author Jasna Kovac, postdoctoral researcher at Cornell's Food Safety Laboratory and Milk Quality Improvement Program. "The whole genome tool gives us better decision-making ability as to whether to recall food contaminated with these bacteria, making it important for consumer safety."
Bacillus is a group of bacteria that appear in all kinds of food. B. cereus may survive pasteurization and other heat treatment processes, posing a challenge for the food industry. Certain isolates of B. cereus create no problems, but other isolates can cause diarrhea, nausea or vomiting when consumed.
Which isolates are which?
Using the whole-genome sequencing tool, food producers can test bacteria found in products or environments to identify the illness-causing isolates.
"This tool becomes useful for food producers as it enables them to identify bacterial isolates that may cause disease and keep the contaminated products from entering the marketplace," Kovac said.
In addition to Kovac, the research "Production of Hemolysin BL by Bacillus Cereus Group Isolates of Dairy Origin is Associated with Whole-Genome Phylogenetic Clade" was conducted by doctoral students Rachel A. Miller, Sarah Beno and Laura Carroll; graduate student Jiahui Jian; technician David J. Kent; and the paper's senior author, Martin Wiedmann, Cornell's Gellert Family Professor in Food Safety.
Whole-genome sequencing is becoming less expensive and Kovac believes it will be used more commonly in combination with traditional methods in the future. "It has the potential to be used throughout the entire food industry," she said. "When food producers need to make a decision fast, this will be their first choice."
In July, the European Food Safety Authority called for applying whole-genome sequencing to characterize and provide detail for bacterial foodborne outbreaks implicating Bacillus. Commenting on the European suggestion, Wiedmann said, "I think this shows a great example of how Cornell research produces data that are immediately needed to make better food safety decisions."
More information: Jasna Kovac et al. Production of hemolysin BL by Bacillus cereus group isolates of dairy origin is associated with whole-genome phylogenetic clade, BMC Genomics (2016). DOI: 10.1186/s12864-016-2883-z