Human 'junk' gene sequences can promote translation
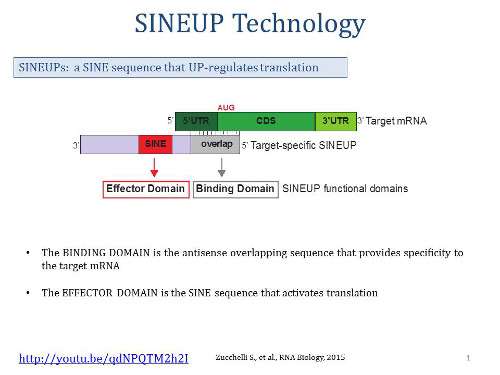
One of the biggest surprises of the past decade of genomic studies was the discovery that, contrary to previous belief, the majority of the genome is not used to produce proteins. Initially, many scientists thought that these long non-coding RNAs were non-functional "noise," but in recent studies, a growing fraction of these lncRNAs have been found to have regulatory functions.
In 2012, a group from the RIKEN Center for Life Science Technologies, in collaboration with SISSA, an Italian University, discovered a new class of mouse lncRNAs, which are called "antisense" because their can pair with typical protein-coding mRNAs and enhance their translation. The team was surprised to find that pairing these novel antisense RNAs caused the mRNA to be translated more efficiently. This was counterintuitive, because it was generally believed that antisense RNAs always inhibited gene activity. They found that the positive effect on specific mRNA translation was mediated by a sequence embedded in the antisense lncRNA. This element turned out to be a member of what is known as the SINE family, standing for "short interspersed elements," which are very broadly distributed throughout the genome. They were long considered to be part of the "junk" of the genome—remnants of ancient parasites in the genome capable of making copies of themselves, filling up the genome with junk.
Specifically, the repeat elements discovered during the earlier research belonged to the SINEB2 class, which is present only in rodent genomes. Artificial RNAs containing these SINEB2 elements, which can be made to target any RNA by simply engineering the antisense region, are called SINEUPs (RNA containing SINE elements that UP-regulate translation). Due to the peculiar evolution of the mouse SINEB2 elements, which are thought to have co-evolved from an ancestor sequence similar to tRNAs, natural SINEUPs were thought to exist only in mice.
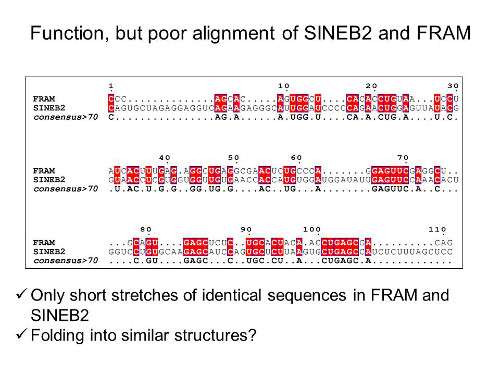
Now, in a discovery reported in Scientific Reports, the group found a group of human sequences—unrelated to those in mice—which were also capable of producing SINEUPs. The group carried out a broad screening of RNA sequences from humans and tested the function of various antisense RNAs to see if they could have SINEUP functionality. Surprisingly, they found that human SINE elements of a very different type from the mouse SINEB2—called FRAM and MIRb—could. According to Piero Carninci of CLST, who led the team, "This was very surprising, because unlike the mouse SINEB2, the human ones resemble unrelated families of non-coding RNAs. We were even more surprised when we looked at the similarity between the mouse and human sequences: the newly discovered FRAM elements share less than 30% of their sequences with the mouse SINEB2 elements, yet they function in a very similar fashion, by enhancing translation of mRNAs they overlap."
"As in the mouse," he continues, "the human elements can be used in biotechnological applications to target mRNA encoding different proteins, by simply engineering the antisense part. Although the primary sequence is so different, the human and the mouse may fold in a similar way and hence have similar function. The structural basis for this functions are yet unknown, and this remains an exciting remaining question."
SINEUPs are currently used in biotechnology applications, to specifically stimulate the translation of mRNAs to produce more proteins both in cell cultures to study gene function and to produce biological proteins in bioreactors. Additionally, laboratories around the world are working to develop SINEUPs to enhance protein translation as a therapy for specific diseases caused by the deficiency of a specific protein, such as haploinsufficiencies, where one of two genes is not functional. According to Carninci, "Our discovery provides one more weapon to the SINEUP toolbox to broaden the scope of gene therapy, as well as to express a greater variety of proteins."
More information: Aleks Schein et al. Identification of antisense long noncoding RNAs that function as SINEUPs in human cells, Scientific Reports (2016). DOI: 10.1038/srep33605