Using DNA to predict schizophrenia and autism
Huntington's disease, cystic fibrosis, and muscular dystrophy are all diseases that can be traced to a single mutation. Diagnosis in asymptomatic patients for these diseases is relatively easy—you have the mutation? Then you are at risk. Complex diseases, on the other hand, do not have a clear mutational footprint. A new multi-institutional study by Japanese researchers shows a potential rare gene mutation that could act as a predictor for two neurodevelopmental disorders, schizophrenia and autism.
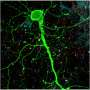
"Aberrant synapse formation is important in the pathogenesis of schizophrenia and autism," says Osaka University Professor Toshihide Yamashita, one of the authors of the study. "Microglia contribute to the structure and function of synapse connectivities."
Microglia are the only cells in the brain that express the receptor CX3CR1. Mutations in this receptor are known to affect synapse connectivity and cause abnormal social behavior in mice. They have also been associated with neuroinflammatory diseases such as multiple sclerosis, but no study has shown a role in neurodevelopment disorders.
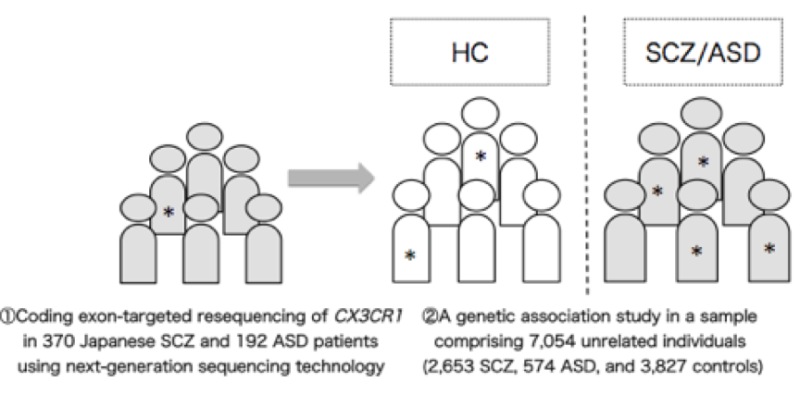
Working with this hypothesis, the researchers conducted a statistical analysis of the CX3CR1 gene in over 7000 schizophrenia and autism patients and healthy subjects, finding one mutant candidate, a single amino acid switch from alanine to threonine, as a candidate marker for prediction.
"Rare variants alter gene function but occur at low frequency in a population. They are of high interest for the study of complex diseases that have no clear mutational cause," said Yamashita, who added the alanine threonine substitution was a rare variant.
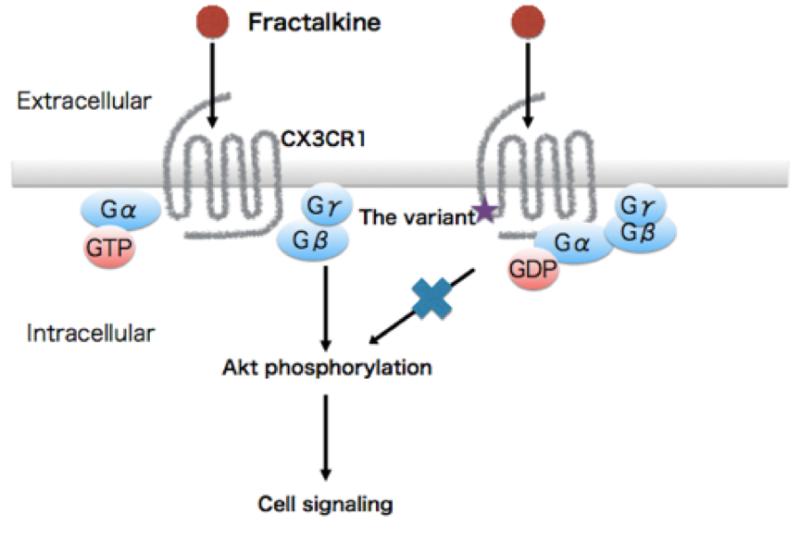
The structure of CX3CR1 includes a domain known as Helix 8, which is important for initiating a signaling cascade. Computer models showed that one amino acid change is enough to compromise the signaling.
"The variant changes the region from hydrophobic to hydrophilic and destabilize Helix 8. We overexpressed the mutation in cells and found Akt signaling was disrupted," explains Yamashita.
According to Yamashita, the findings are the first to connect a genetic variation in microglia with neurodevelopmental disorders. Moreover, he hopes that the discovery could become a basis for predictive diagnostics.
"There is no reliable way to diagnose schizophrenia or autism in asymptomatic patients. Deeper understanding of the genetic risk factors will help us develop preventative measures."
More information: K Ishizuka et al, Rare genetic variants in CX3CR1 and their contribution to the increased risk of schizophrenia and autism spectrum disorders, Translational Psychiatry (2017). DOI: 10.1038/tp.2017.173