Researchers discover how enzyme 'shape-shifts' in drug-resistant leukemia

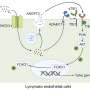
St. Jude Children's Research Hospital structural biologists have deciphered how the structure of the enzyme called Abl regulates its activity, enabling the enzyme to switch itself on and off. Understanding Abl's regulation is important because a mutant form of the enzyme (Bcr-Abl) is over activated in chronic myelogenous leukemia and other cancers. Abl is a central growth-controlling switch in white blood cells. The enzyme's over activation spurs mutated cells to the uncontrolled growth of leukemia.
While clinicians have had success in treating CML with drugs that switch off the enzyme, it often mutates to become drug resistant. The new study reveals the aberrant mechanism underlying some drug-resistance mutations that have remained mysterious. The study's findings also offer a path to possible treatments to overcome resistance.
The research was led by Charalampos Kalodimos, Ph.D., chair of the St. Jude Department of Structural Biology, and appears today as an advance online publication in the journal Nature Structural & Molecular Biology.
In their experiments, the researchers sought to understand how Abl manages to switch itself on and off by altering its shape. The Abl enzyme controls this switching through a process called allosteric regulation in which a part of the molecule distant from the molecule's on-off switch or kinase domain somehow inhibits or activates Abl.
"We knew we had these two functional states, but we had no idea about the conditions under which Abl switched from one to another," Kalodimos said. "We also didn't understand how external molecules that regulate Abl acted on these two states. Nor did we understand how mutations that confer drug resistance affected the states."
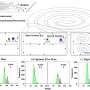
To probe the Abl molecule's constant shape-shifting, the researchers used an analytical technique called NMR spectroscopy, in which strong magnetic fields are used to tweak a molecule to produce signals that reveal its structure. NMR spectroscopy can uniquely "see" molecules undergoing rapid changes. The researchers' experiments explored in detail how the region of Abl called the allosteric regulatory module interacted with the kinase domain to control it.
Their experiments revealed that in its shape-shifting, Abl was precisely balanced between its inhibition and activation states, Kalodimos said.
"We saw this very fast 'breathing' motion of several thousand times a second, in which the molecule goes on and off, on and off," he said. "This motion is important, because it allows other molecules that regulate Abl to adjust its activity one way or the other in a graded manner—like turning a rheostat up or down." Such regulation would involve pushing the Abl molecule toward either the inhibited or activated shape, he said.
The fact that NMR spectroscopy captures the complex conformation changes of molecules enabled the researchers to discover new details about how Abl's structure affected its activation state. For example, their experiments revealed a previously unknown activator region embedded in the molecule. This region is likely responsible for one mutation that produces drug resistance, Kalodimos said.
The researchers also used their technique to analyze mutations in Bcr-Abl that enable it to become resistant to the drug imatinib, known as Gleevec. The drug has proven effective in treating CML by plugging into the kinase domain of the over-activated Abl enzyme and shutting it down. However, in many patients a mutation in the gene that produces Abl renders it drug resistant. While many of the mutations block Gleevec from plugging into the kinase domain, others appear to interfere with the allosteric regulation. In effect, they may "warp" the enzyme to keep it activated.
In analyzing the structure of these allosteric mutants, Kalodimos and his colleagues discovered that they altered Abl's shape to activate it; and did not interfere with how Gleevec plugs into the kinase domain. This finding points the way to new treatments to overcome such resistance, Kalodimos said.
"There is now a new generation of drugs that bind to the allosteric pocket to inhibit its activity," he said. "These could be combined with Gleevec to overcome allosteric mutations to shift Abl into an inhibited state."
Kalodimos said that the treatment strategy could also be applied to other forms of leukemia that have uncontrolled Bcr-Abl activity. The new basic understanding of Abl regulation will also yield insight into similar enzymes, in which allosteric regulation controls a kinase domain.
More information: Atomic view of the energy landscape in the allosteric regulation of Abl kinase. Nature Structural & Molecular Biology. DOI: 10.1038/nsmb.3470