New assay may boost targeted treatment of non-Hodgkin lymphoma
Diffuse large B-cell lymphoma (DLBCL) is an aggressive cancer and the most frequently diagnosed non-Hodgkin lymphoma worldwide (nearly 40% of cases). Recent advancements indicate that both the prognosis and choice of treatment of DLBCL may depend on identifying its molecular subtype. In a report in the Journal of Molecular Diagnostics, researchers describe their development of a reliable, accessible, rapid, and cost-effective new gene expression signature assay that can enhance lymphoma management by helping to match tumors with the appropriate targeted therapy.
"DLBCLs are heterogeneous, with three major subtypes associated with different outcomes. According to the recent update of the World Health Organization (WHO) classification, this information should therefore be systematically provided at diagnosis. Moreover, many targeted therapies are under clinical evaluation and being able to distinguish these diseases routinely and accurately should soon become a major goal," explained lead author Victor Bobée, PharmD, of the Henri Becquerel Cancer Treatment Center, INSERM U1245 (Rouen, France).
For example, the germinal center B-cell-like subtype (GCB) has a favorable prognosis, with a five-year overall survival rate of almost 75%. In contrast, the subtype known as activated B-cell-like (ABC) is more aggressive and less responsive to current therapies but may respond to NF-kB pathway inhibitors, such as ibrutinib. "Accurate diagnostic methods capable of discriminating these subtypes are thus needed," said co-investigator Philippe Ruminy, PhD, of the Henri Becquerel Cancer Treatment Center, INSERM U1245 (Rouen, France).
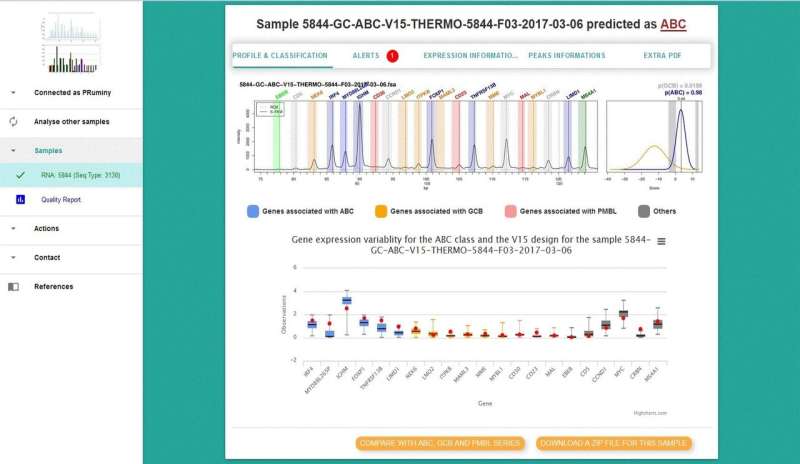
The described assay is a novel gene expression profiling DLBCL classifier based on reverse transcriptase multiplex ligation-dependent probe amplification (RT-MLPA). It has the capability to simultaneously evaluate the expression of 21 markers, allowing differentiation of the three subtypes of DLBCLs (GCB, ABC, and primary mediastinal B-cell lymphoma, PMBL) as well as other individualized disease characteristics, such as Epstein Barr infection status.
One hundred fifty RNA samples extracted from biopsies were tested. Forty-two percent of the samples had the ABC subtype, 37% the GCB subtype, and 10% molecular PMBL. Eleven percent of samples could not be classified. Overall, the RT-MLPA assay correctly assigned 85.0% of the cases into the expected subtypes compared to 78.8% with immunohistochemistry.
The assay was also able to detect the MYD88 L265P mutation, one of the most common genetic abnormalities found in ABC DLBCLs. This information can influence treatment, since the presence of the mutation has been suggested to be predictive of ibrutinib sensitivity.
Currently, in most institutions the identification of the different DLBCL subtypes at diagnosis is addressed through immunohistochemistry, which often relies on the skills of experienced hemato-pathologists with expertise in the field of lymphoma.
RT-MLPA is a robust, efficient, rapid, and cost-effective alternative to the current methods used in the clinic to establish the cell-of-origin classification of DLBCLs. In contrast to other technological approaches such as RNA sequencing or nanostring technologies, its implementation requires only common laboratory equipment and not dedicated and expensive platforms. Another advantage is that it can be applied to formalin-fixed, paraffin-embedded samples. Other types of diagnostic methods may not provide the level of detail needed and may also be limited by poor reproducibility and lack of adaptability to routine use in standard laboratories.
"Because we have provided the classification algorithms, other laboratories will be able to verify our results and adjust the procedures to suit their environment," noted Dr. Ruminy. "It is our hope that the assay we have developed, which addresses an important recommendation of the recent WHO classifications, will contribute to better management of these tumors and improved patient outcomes."
More information: Journal of Molecular Diagnostics (2017). DOI: 10.1016/j.jmoldx.2017.07.007