Big Data shows how cancer interacts with its surroundings
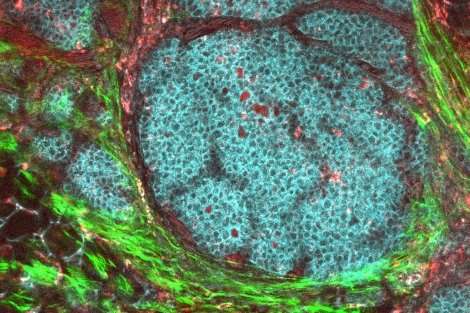
By combining data from sources that at first seemed to be incompatible, UC San Francisco researchers have identified a molecular signature in tissue adjacent to tumors in eight of the most common cancers that suggests they are all using the same mechanism to remodel normal tissue and spread.
The new study is the first systematic analysis of the normal-looking tissue near tumors that gets removed in cancer operations. Precision medicine researchers use this so-called normal adjacent tissue, which looks normal under a microscope and is usually at least two centimeters from tumors, as a basis of comparison to highlight the changes that occur in cancer. But the new study suggests the tissue is far from normal at the molecular level and is rather somewhere in between cancerous and healthy. The analysis demonstrates how tumors of many different types may be instigating inflammatory and other cancer-related processes in other tissues, to facilitate their spread.
"Tumors secrete factors all around, changing nearby tissue and possibly even tissues that are far away," said Dvir Aran, PhD, a postdoctoral fellow at the UCSF Institute for Computational Health Sciences (ICHS) and the first author of the paper, published Oct. 20, 2017, in Nature Communications. "We saw more or less the same effects across all the major cancer types, which suggests this is an important mechanism for the tumor."
Much of what the study found actually replicates the findings of laboratory studies of how various types of cancer interact with their surroundings. What it adds, however, is a comprehensive view of how all the different cancers are using similar strategies to alter tissue outside their boundaries.
"The whole cancer world is focused on trying to figure out what the environment of these cancer cells is really like," said ICHS Director Atul Butte, MD, PhD, who is the Priscilla Chan and Mark Zuckerberg Distinguished Professor in the UCSF Department of Pediatrics and also the senior author of the paper. "We've got to do more work like this to understand how the cancers are growing and thriving."
The researchers used data from The Cancer Genome Atlas (TCGA) to analyze which genes were being turned on or off in the tissue adjacent to tumors in eight major cancers: lung, colon, breast, uterine, liver, bladder, prostate and thyroid. They found that genes involved in the acute phase of systemic inflammation had been turned on, creating a pattern that was distinct from both the tumors and the healthy tissue from similar spots in the body found in another database – the Genotype-Tissue Expression (GTEx) program – that they used as a comparison. The GTEx program collected data from many different places in the body in hundreds of patients who died in the hospital and donated their bodies for research. It excluded tissue from donors who died of cancer or who had received chemotherapy or radiation within two years, so while the samples may not be completely healthy, they provided a good contrast to the cancerous and tumor adjacent tissue in TCGA.
Comparing the two databases presented several technical challenges. First, the researchers had to parse out the statistical variation that comes from slight differences in computational methods, then they had to find a way to control for the "batch effects" that come when data is collected at different times and places by different people. This was particularly hard to do, since there was no overlap between the two databases. But they found a way to use a statistical technique developed for RNA sequencing data called "remove unwanted variation," and wound up with a combined database with information on 1,558 normal control samples, 428 adjacent tissue samples, and 4,500 cancer samples.
To demonstrate the validity of their initial finding that the tumor adjacent tissue was different from both cancerous and healthy tissue, the researchers analyzed data from smaller public repositories with adjacent and tumor samples from colon, liver, breast and prostate cancers, along with healthy tissue samples from the corresponding locations in the body. These samples are not from combined projects, and are therefore less affected by the statistical noise the researchers were trying to remove from their larger assembled dataset. The data in the smaller datasets broke cleanly between the three tissue types in colon, liver and breast cancer, and trended that way in prostate cancer.
This finding echoed a pattern the researchers saw in their big dataset: the tissue adjacent to cancers with more distinct boundaries, like breast, colon, liver, lung and uterine cancer, was more clearly defined by the data than the tissue adjacent to cancers of the prostate and some types of thyroid cancer, in which tumors are more diffuse.
The big data analysis identified 82 genes that were upregulated in the adjacent tissue, compared to both healthy and cancerous tissue. In general, the researchers found the adjacent tissue more closely resembled normal tissue than cancerous tissue when it came to the activation of certain hallmark cancer-related genes like MYC. Adjacent tissue was in between cancerous and normal in how its fat and muscle cells grew. And it looked cancerous when it came to inflammation.
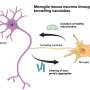
An inflammatory signaling pathway, possibly regulated by tumor necrosis factor-alpha, which is involved in the acute phase of inflammation, was highly expressed in seven out of eight of the adjacent tissue types analyzed. The adjacent tissue also had more immune cells than normal tissues do, supporting the idea that inflammation was at work.
The researchers found the tumors were most likely acting on the blood vessels of the surrounding tissue, to remodel it with processes the tumor had already used during its own formation. The researchers also found evidence that the adjacent tissue was under the influence of cancer-related stress signals that induce oxygen deprivation and cell death. And they saw other notorious cancer processes at work, including one that causes mature cells to return to an embryonic state, from which they can proliferate into cancer cells.
"Cancer research is focused on the tumor microenvironment – how the tumor is changing its own environment – but there are changes that go beyond that, which we should also look at," Aran said.
The researchers confirmed their findings about how the tumors were remodeling nearby tissue by reanalyzing data from a smaller study of how tumors change surrounding breast tissue. Then they did an experiment in mice, using a gene they had identified in the big data study. They implanted human breast cancer tissue into the mice, then measured the mouse equivalent of that gene's expression level. To their surprise, they found it was activated not only in the mammary gland with the cancerous implant, but also in the opposing one. Looking further in human breast tumors they found that gene was being expressed particularly in endothelial cells, which line the interior of blood vessels. This suggests the cancer was emitting a signal to make blood vessels grow in nearby tissue.
The finding was one of several in the paper that could be pursued as a therapeutic target.
"We could try to block it and see if the tumor is still able to spread," Aran said. "If you block this pathway, the tumor can't get what it needs from the blood vessel."
Overall, the researchers said their study suggests that precision-medicine researchers should be cautious about drawing too many conclusions from datasets that do not include truly normal tissue, since comparing tissue that has already been partially remodeled by cancer inevitably understates just how different tumors are from healthy tissue.
"When we began this experiment, we wanted to know if it was problematic to compare cancer to seemingly normal adjacent tissue samples," Aran said. "After analyzing the data, we found that by understanding the differences between truly healthy and tumor adjacent tissues, we were actually learning interesting things about cancer."
More information: Dvir Aran et al. Comprehensive analysis of normal adjacent to tumor transcriptomes, Nature Communications (2017). DOI: 10.1038/s41467-017-01027-z