Three-dimensional folding of DNA provides important epigenetic mechanisms in the formation of cardiac muscle cells
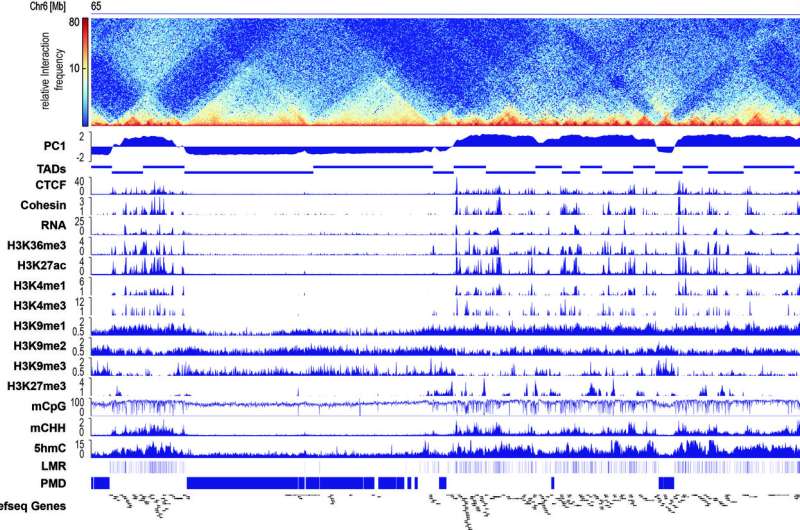
During differentiation of pluripotent stem cells to cardiomyocytes, the three-dimensional folding of the DNA reorganizes itself. This reorganization of the DNA architecture precedes and defines important epigenetic patterns. A team lead by private lecturer Dr. Ralf Gilsbach and Stephan Nothjunge, who both conduct research at the University of Freiburg in the Department of Experimental and Clinical Pharmacology and Toxicology headed by Prof. Dr. Lutz Hein, have come to this conclusion. The results suggest that the genome's spatial organization is an important switch for defining cell types, thereby representing a very promising starting point for future reprogramming strategies. The team recently published its results in the scientific journal Nature Communications.
The genome stores information about an organisms development. Each cell carries this information tightly packed on a two-meter long DNA strand in the cell nucleus and specific epigenetic mechanisms control access to the 'blueprint of life'. Because every cell type in a mammalian organism requires access to genomic areas in a tempo-spatial specific manner, the epigenome is crucial for determining cellular identity. It is already known that various epigenetic mechanisms are associated with cell differentiation. Particularly indispensable is the methylation of DNA, in which methyl groups are attached to specific nucleotides of double-stranded DNA. Recent studies also show that differentiation processes are accompanied by a reorganization of the three-dimensional folding of the DNA. Up until now, however, it has been unclear what comes first during cardiomyocyte differentiation: the reorganization of the DNA's folding in the cellular nucleus or the DNA's methylation - and whether these mechanisms are dependent on one another.
In order to address this question, the team lead by the Freiburg pharmacologists used modern sequencing methods. These made it possible to map the three-dimensional genome organization as well as epigenetic mechanisms during the differentiation of cardiomyocytes across the entire genome. For this purpose, the researchers established methods for isolating cardiomyocytes in various developmental stages from healthy mouse hearts. This cell-type-specific analysis was essential to demonstrate that there is a close interplay between epigenetic mechanisms and the spatial folding of the DNA in the cardiomyocytes' nucleus. The comparison of different stages of development showed that the type of spatial folding of DNA defines which methylation patterns are formed and which genes are activated. The researchers proved that the spatial arrangement of the DNA is not dependent on the DNA methylation with cells, among other things, that have no DNA methylation at all. The three-dimensional genome organization is thus a central switchboard for determining cellular identity. In the future, the researchers want to use this switch to control cellular functions.
More information: Stephan Nothjunge et al, DNA methylation signatures follow preformed chromatin compartments in cardiac myocytes, Nature Communications (2017). DOI: 10.1038/s41467-017-01724-9