Scientists unlock structure of mTOR, a key cancer cell signaling protein
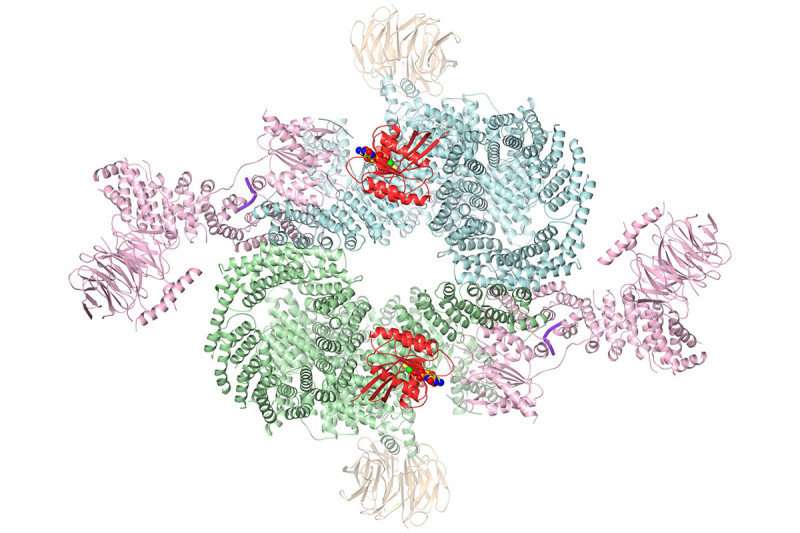
Researchers in the Sloan Kettering Institute have solved the structure of an important signaling molecule in cancer cells. They used a new technology called cryo-EM to visualize the structure in three dimensions. The detailed information provided by this image paves the way for rational drug design.
Scientists call it the master growth regulator: a protein complex in cells that senses when they have enough nutrients to grow and divide.
When this complex, called mTOR, gets triggered, cells begin making copies of key ingredients, such as membranes, DNA, and organelles. They'll need these extra materials to divvy up when they split into daughter cells.
Cancer cells need mTOR too. In fact, many cancers have found ways to keep the mTOR protein active essentially all the time. That's part of the way they ensure limitless cell division.
For years, scientists have sought to target abnormally active mTOR with drugs as a way to treat cancer. Two such drugs are FDA approved, for the treatment of some types of kidney and breast cancer.
But overall, mTOR-targeted drugs have been disappointing. That may be because mTOR is a large and complex piece of cellular equipment. There are many interacting parts, and it may be hard to take down the whole thing with a single shot.
Now, a team of scientists at the Sloan Kettering Institute (SKI) led by structural biologist Nikola Pavletich has assembled an impressively detailed view of the protein complex, including what it looks like in action. This ultra-high-resolution picture could lead to the design of more-effective drugs to block this key cancer driver.
Freeze Frame: Cryo-EM Captures Complete Picture of mTOR
The new picture is the product of a cutting-edge research tool called cryo-electron microscopy (cryo-EM). This tool—which won its developers this year's Nobel Prize in Chemistry—shoots beams of electrons at proteins that have been flash frozen in liquid nitrogen. The electrons bounce off the sample in precise ways, forming an image. The process isn't so different from the way a camera works: light bounces off an object and into your camera lens, producing a photographic image. With cryo-EM, thousands of such pictures are taken, and then a computer reassembles them into a precise, three-dimensional image.
Previously, such detailed atom-by-atom pictures could only be produced with X-ray crystallography, a painstaking and time-consuming method that involves first making a crystal out of a protein. But not all proteins will form crystals. That's especially true of large proteins and protein complexes with multiple moving parts—like mTOR.
Cryo-EM eliminates the crystallization step. This makes determining structures with this technique is much easier and faster.
"We actually had tens of milligrams of this protein back in 2008, but we couldn't crystallize the whole thing," says Haijuan Yang, a senior research scientist in the Pavletich lab at SKI who is the first author on a new paper published in Nature.
So instead, she and her colleagues started truncating the protein, lopping off bits until they got a portion that would crystallize. They published their findings of this smaller fragment in 2013.
Like many proteins, mTOR is an enzyme, which binds to target molecules called substrates in a precise way to hasten chemical reactions. Specifically, it is a type of kinase, which removes phosphates (P) from ATP, the cell's energy currency, and places them on other molecules. From that 2013 study, which took more than five years to complete, the team learned some key things about the protein, such as what the ATP-binding site looks like and where a drug called rapamycin binds.
But it was still only a partial view. The mTOR protein is actually part of a larger assembly of several interlocking protein subunits that operate together. The whole complex is called mTORC1. In addition to the subunit that binds ATP, there is also a subunit called RAPTOR that interacts with other proteins as part of a signaling chain. The mTOR enzyme is activated by a protein called RHEB and inhibited by one called PRAS40.
The structure newly obtained by cryo-EM shows how all these pieces fit together, including how the mTOR enzyme is turned on.
"We figured out that RHEB causes a major conformational change in the complex," Dr. Yang says. "RHEB binding creates a new interface that doesn't exist without this binding, then twists the mTOR kinase domain to bring it into the active conformation."
Finally, they showed how common cancer-associated mutations that occur in this protein effectively keep it permanently in the active state.
Latest in a Long Line of Cancer-Related Structures
Solving the structure of mTORC1 is just the latest in a long line of protein structures that Dr. Pavletich and members of his lab have solved. In the 1990s, they deciphered the structure of p53, the most commonly mutated protein in cancer. And in the early 2000s, they determined the structure of BRCA2, a tumor-suppressor protein involved in DNA repair. When mutated, this protein is associated with inherited predispositions to breast cancer, ovarian cancer, and prostate cancer.
Dr. Yang says they plan to return to the BRCA2 protein. Studying it with cryo-EM will allow them to obtain even more information than was previously possible.
"Before, people could model bits of this protein and bits of that protein," she says. "Now, with cryo-EM, we can put it all together."
More information: Haijuan Yang et al. Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40, Nature (2017). DOI: 10.1038/nature25023