Natural lung bacteria offer clues on treating airway infections, says expert
Further research on the lung microbiome could uncover new, more effective approaches to managing lung infections, according to an Imperial researcher.
An Imperial College London expert has highlighted the importance of studying lung microorganisms, known as the lung microbiome, in helping us to understand and manage airway infections.
Professor William Cookson, from Imperial's National Heart and Lung Institute, spoke to Francesca Davenport about his Nature Reviews Microbiology paper, to explain how further studying the microbiome could help to improve the management of lung infections and avoid problems associated with antimicrobial resistance (AMR).
What were your reasons for writing the review paper?
Lung infections are amongst the most important causes of death and disability in the world, but progress in diagnosing and treating them has lagged far behind infections of other organs such as the gut and skin. In particular, we have only just begun to use DNA sequencing to identify the bacteria and funguses that we all carry in our airways.
Our article highlights the size of this problem, and shows how modern microbiome research has the potential to improve the prevention and treatment of common lung diseases.
How much do we currently know about the lung microbiome?
Until recently, scientists incorrectly believed that the lungs and airways were sterile, and so the Human Microbiome Project and similar research ventures almost completely ignored the lungs.
Now we know the surfaces of the airways have a resident population of microbes, known as the microbiome, which is similar in density to that of the skin and upper intestines. We also know these microbial communities are disturbed in common lung diseases like chronic obstructive pulmonary disease (COPD), which is common in smokers, and asthma.
However, other than the few organisms that commonly cause lung infections, we know surprisingly little about the different bacteria that live there. We don't yet know how members of the microbial community interact with each other, and how they might control inflammation and protect the airway linings from disease.
What are the specific challenges to researching the lung microbiome?
It is more difficult to get samples from the airways than easier to access areas like the skin and bowel. Subjects can cough up mucus for testing, or clinicians can introduce a fine tube, or bronchoscope, into the airways.
Difficulties arise because lung samples contain many human cells, and the presence of high concentrations of human DNA means special techniques are necessary for the bacterial signals to be seen.
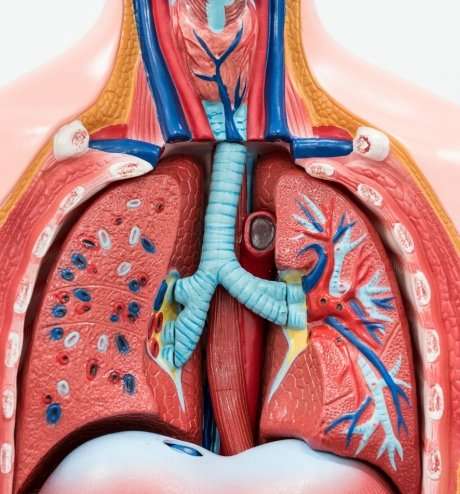
How do we develop and maintain a balanced and healthy lung microbiome?
There are one or two hundred different kinds of bacteria in the lungs of healthy individuals. Taken across the population there are several thousand different bacterial strains.
Normal colonisation of the airways likely begins after birth, and many of the organisms may initially come from the mother. Over the first two years of life, the microbiome becomes more complex and species-diverse. As with other natural ecosystems, it is likely that it is the diversity of the microbiota in the lungs protects them against infection with harmful miroorganisms.
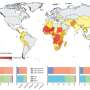
A rural lifestyle, for example living on farms, encourages diversity for the microbiome in the lungs as well as the bowel and skin. We have known for some time that rural living protects against asthma and allergies. Living in cities decreases the species-diversity, and also provides many more opportunities for harmful bacteria to spread in the community.
What are the problems with traditional diagnostics and treatments of lung infections?
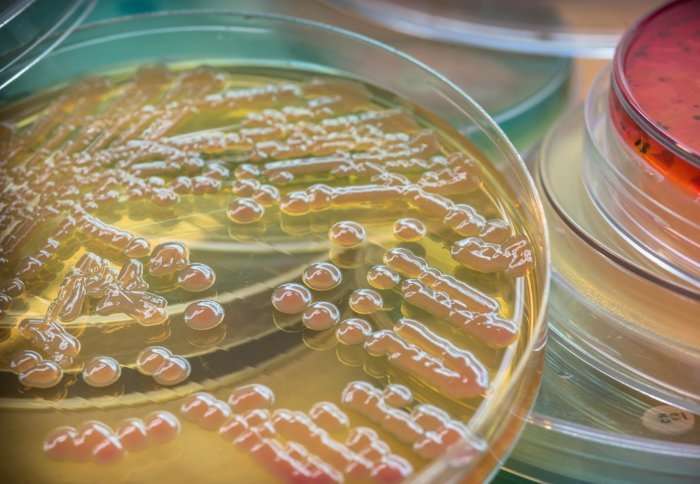
In our society, lung infections most often arise in people with pre-existing lung diseases such as COPD and diseases such as asthma also show marked abnormalities in their airway microbiome. Infections are usually diagnosed by culturing the bacteria, where specimens from patients are spread on agar plates and incubated for one or more days until the bacteria can be seen as colonies on the plate. However, it is important to treat lung infections more quickly than the timeframe allowed by culturing.
Because culturing takes so long, doctors often have to guess at the cause of infections and prescribe broad spectrum antibiotics that wipe out both good and harmful bacteria. As a result, there are approximately 16 million antibiotic prescriptions for respiratory infections each year in the UK; equivalent to a quarter of the population. Thus the timing issue is a major cause of AMR in our society.
What improvements have there been in diagnostics with modern sequencing? How can these improve treatments and avoid problems with AMR?
Once we are sure exactly which organisms cause lung disease, we can use rapid sequencing methods to find if these bacteria are present.
We will also be able to estimate their numbers, tell how dangerous they are, and to which antibiotics they will respond. With improved diagnosis, we can target antibiotics towards the bad bacteria and do less damage to good bacteria. We can also make antibiotics for inhaling directly into the lungs, which will protect against any side effects on other body systems.
Another new approach is to use bacteriophages, which are very small viruses that can infect and destroy bacteria. They may be used to treat infections in conditions such as cystic fibrosis, where almost continuous antibiotic therapy has led to the emergence of 'super-pathogens' that are resistant to almost all existing antibiotics.
We will eventually be able to to replace abnormal airway bacteria with a healthy population of good bacteria, similar to planting a meadow in a field that has previously been stripped of vegetation. Using good bacteria to recolonise the microbiome is hugely effective in treating bowel conditions, and the same is likely to be true for lung and airway diseases.
In the paper you suggest developing an international lung microbiome project – can you describe how this would work?
Only a few of the thousands of organisms that can be found in normal airways have been isolated and studied in any detail. For sequence-based diagnostics to be effective, we need to know as much as possible about the good and bad bacteria in the lungs. International collaborations to isolate and sequence bacterial DNA have hugely improved the diagnosis and treatment of infections in the mouth, skin and bowel. A concerted lung microbiome project will speed up the development of all the potential advances in diagnosis and therapy described in our review.
More information: William O. C. M. Cookson et al. New opportunities for managing acute and chronic lung infections, Nature Reviews Microbiology (2017). DOI: 10.1038/nrmicro.2017.122