Researchers develop technique to analyse cancer cells' life history
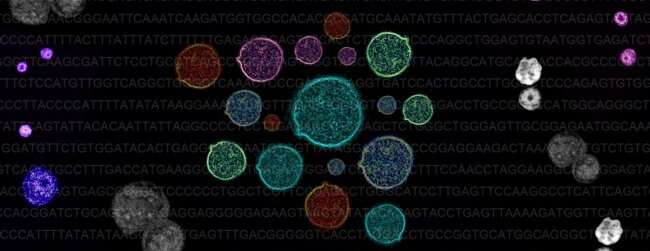
A team of researchers from the MRC Weatherall Institute of Molecular Medicine at Oxford University has developed a new technique that allows scientists to reliably track genetic errors in individual cancer cells, and find out how these might lead to uncontrollable growth.
Despite recognising that cancer cell diversity underlies treatment resistance and recurrence of cancer, previous attempts to track errors in individual cancer cells were very inaccurate, or could only track a few cells at a time.
This is the first time that researchers have been able to reliably track DNA errors, or 'mutations' in thousands of individual cancer cells, while also measuring how these mutations lead to disruption to how DNA is read within individual cancer cells in a tumour.
The study, published in the journal Molecular Cell, describes how this new technique, TARGET-seq, can not only detect mutations within individual cancer cells from patients, but also work out the full list of gene products in individual cancer cells (the transcriptome). Tracking these genetic errors, and their consequences, is important because despite the latest medical advances, completely getting rid of cancer cells is sometimes extremely difficult. As there are many different kinds of cancer cells in a tumour, they can all behave differently and have different kinds of resistance to treatment. Understanding the genetics of individual cancer cells in such detail will help clinicians personalise cancer treatments for each patient.
Professor Adam Mead, a MRC Weatherall Institute of Molecular Medicine researcher who also works as consultant NHS physician treating blood cancer patients, said: "Without knowing what kinds of cells a patient has, it is difficult to predict how the patient will respond to a particular kind of treatment or drug. This means that cancer patients often have relapses, because their treatment only killed off some kinds of cancer cells".
Professor Mead's team at the MRC Molecular Haematology Unit in Oxford University's Radcliffe Department of Medicine used their TARGET-seq technique on tumour samples from 15 patients whose blood-making cells had become cancerous.
This is the first time that researchers have directly tested individual cancer cells to find that even cells from the same patient, with the same type of blood cancer, are quite different from each other, depending on the genetic errors that each of them has accumulated.
The pattern of genome and transcriptome errors was so detailed that the researchers could reconstruct the 'life history' of every single cancer cell in each tumour, uncovering the order in which the genetic mutations in each tumour cell happened.
Surprisingly, even patients that had very similar cancer and had been treated with the same therapies had very different tumour 'life histories'.
Professor Mead's team now hope to apply their technique to hundreds of patient samples, to be able to identify the most common kinds of events and sequences that take place in cancer cells' 'life history'.
"Having this information is really important, because it not only gives us unique information about how tumours change over time, but also how they might respond in future to different treatments and drugs." said Alba Rodriguez-Meira from the MRC Weatherall Institute of Molecular Medicine, the first author on the study.
"No two patients will have exactly the same mixture of cancer cells with exactly the same pattern of evolution, and our technique will allow clinicians to monitor patient progression during clinical trials and ultimately customise the treatment they offer to the unique mixture of cancer cells that every cancer patient has."
Professor Mead, added: "This technique will ultimately help us track very rare subpopulations of cancer cells that are resistant to current treatments. Understanding exactly how cancer cells develop also means that we can design new therapies from the ground up, which specifically target the early stages of this development so that we can assess whether a certain treatment is working or develop new rational therapeutic approaches to be able to completely eliminate tumour cells."
Dr. Mariana Delfino-Machin, Programme Manager for Cancer at the MRC, added: "Cancer is often thought of as a single disease, but it is actually a collective term for the abnormal growth and spread of cells, which can differ even within the same patient. If we are to treat cancer successfully, we need to understand exactly how all these different abnormalities behave and respond to treatment, and this is hugely challenging. This new technique takes us one step closer to achieving this goal. If we can understand the background and behaviour of individual cancer cells in a tumour, we have the potential to tailor more effective treatments to the individual in the future."
More information: Application of Single Cell Genomics to Reconstruct Stem and Progenitor Cell Hierarchies in Normal and Malignant Haematopoiesis. gtr.ukri.org/projects?ref=MC_UU_12009%2F16