Predicting cancer versus autism risk in PTEN patients

In a new study published in American Journal of Human Genetics, a team of researchers led by Charis Eng, M.D., Ph.D., Chair of Cleveland Clinic's Genomic Medicine Institute, identified a metabolite that may predict whether individuals with PTEN mutations will develop cancer or autism spectrum disorder (ASD).
Germline mutations of the tumor suppressor gene PTEN are associated with a spectrum of rare genetic disorders that increase the risk of certain cancers, cognitive and behavioral deficits, benign growths and tumors (i.e., hamartomas), and macrocephaly. These disorders are referred to collectively as PTEN hamartoma tumor syndrome (PHTS), but clinical manifestations vary greatly among patients and often are difficult to anticipate.
For example, subsets of Cowden syndrome (CS) and Bannayan-Riley-Ruvalcaba syndrome (BRRS), two well-defined disorders on the PHTS spectrum, are characterized by either a high risk of certain cancers or ASD. There are functional and structural differences between PTEN mutations associated with ASD and those associated with cancer. However, a biomarker that could proactively determine if a patient with CS/BRRS will develop cancer or ASD has not yet been identified.
Previous studies have established metabolic dysregulation as one of the hallmarks of cancer. Specifically, germline variants in the SDHx genes cause an accumulation of the metabolite succinate, which has been linked to tumorigenesis. Some patients with PTEN mutations have been found to have succinate accumulation despite the lack of SDHx mutations, suggesting that variations in metabolite levels may indicate susceptibility to cancer versus ASD.
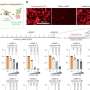
To investigate this further, Dr. Eng's team analyzed the metabolite levels of 511 patients with CS, BRRS, or Cowden-like syndrome compared to controls. The results suggest that certain metabolites are associated with specific mutations and/or clinical features.
In particular, they discovered that decreased levels of fumarate, a metabolite formed from succinate, was more strongly associated with ASD or other developmental disorders compared to cancer in individuals with PTEN mutations. These findings indicate that certain metabolites, such as fumarate, may serve as predictive biomarkers that could distinguish patients who will develop neurodevelopmental disorders from those who will develop cancer.
"By identifying a way to differentiate those with germline PTEN mutations who develop cancer and those who develop autism, this provides clinicians with a new tool to better tailor treatments to individual patients," says Dr. Eng.
Dr. Eng is also the inaugural Director of the Center for Personalized Genetic Healthcare and Medical Director of the PTEN Multidisciplinary Clinic. Notably, she was the first to discover a link between mutations in PTEN and Cowden and other syndromes, and her research has formed the basis of national practice guidelines for those with PHTS and Cowden syndrome. Dr. Eng holds the Sondra J. and Stephen R. Hardis Endowed Chair in Cancer Genomic Medicine at Cleveland Clinic.
More information: American Journal of Human Genetics (2019). www.cell.com/ajhg/fulltext/S0002-9297(19)30343-X