New cardiac fibrosis study identifies key proteins that translate into heart disease
Using cutting-edge technologies, researchers at Duke-NUS Medical School, Singapore, have developed the first genome-wide dataset on protein translation during fibroblast activation, revealing a network of RNA-binding proteins (RBPs) that play a key role in the formation of disease-causing fibrous tissue in the heart. Their findings, published in the journal Circulation, could help in the search for treatments for this condition.
Cardiac fibrosis, a condition characterized by scarring in the heart, is caused by the activation of fiber-producing cells called fibroblasts—which form one of the largest groups of cells in the heart—and underlies many heart diseases, including atrial fibrillation, dilated cardiomyopathy, and heart failure. This involves the transformation of fibroblasts into myofibroblasts, which leads to thickening and stiffening of the heart wall, making it less contractile and thus less able to pump blood around the body.
"Heart disease is a prominent cause of mortality, accounting for one in three deaths in Singapore. In most cases, they are preceded by the transition of resident fibroblasts to myofibroblasts," explained computational geneticist Dr. Owen Rackham, corresponding author of the study and Assistant Professor in the Cardiovascular and Metabolic Disorders (CVMD) Programme at Duke-NUS. "Despite the serious risk and high prevalence of cardiac fibrosis, existing therapies are ineffective and there is an unmet need for new therapeutic approaches to prevent, limit, or reverse the condition."
Being able to disentangle the processes underlying the transformation of fibroblasts into myofibroblasts could help unveil novel molecular pathways underpinning disease onset and pathophysiology, and aid in the search for novel therapeutic targets. The team of researchers from Duke-NUS and colleagues in Germany and the UK investigated the processes that regulate the transcription of DNA code into RNA, and the translation of that code from RNA for protein synthesis during the transformation of fibroblasts into myofibroblasts.
"We found a staggering one-third of all genes undergo translational regulation during this pathogenic transition," highlighted Ms Sonia Chothani, first author of the study and a Ph.D. student at Duke-NUS. "All these gene expression changes are missed or misinterpreted in traditional RNA-based studies."
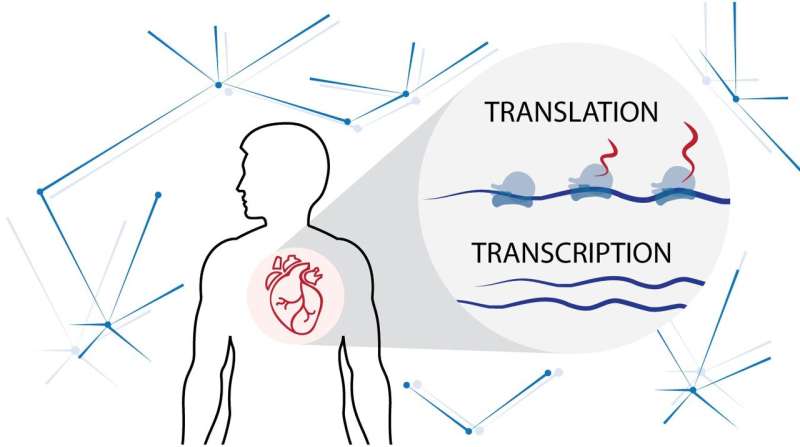
The team first analyzed the gene changes that occurred during DNA transcription and RNA translation at different time points in the fibroblast-to-myofibroblast transition. A computational analysis of this data identified specific regulatory processes affecting RNA translation. They then analyzed RNA found in fibroblasts from tissue samples taken from patients with dilated cardiomyopathy. Many of the regulatory processes identified in the computational analysis were active in the diseased tissue samples.
Specifically, the researchers found RBPs play critical roles in the fibroblast-to-myofibroblast transformation. RBPs target RNA, affecting the translation of its code during protein synthesis. Inhibiting two of these RBPs, called PUM2 and QKI, limited the transformation of fibroblasts into myofibroblasts.
"There are more than 1,500 RBPs encoded in the human genome, but their role in the regulation of translation of target messenger RNAs remains largely unexplored. Our findings show the central importance of translational control in fibrosis, and highlight novel pathogenic mechanisms in heart failure," said Dr. Rackham. "Just as transcription factors are emerging targets in pharmacology owing to their centrality in the dysregulation of transcription, we show that RBPs may play a similar role in the dysregulation of translation."
"Our study combines the use of primary cardiac fibroblasts and heart tissue samples with cutting-edge technologies, leading to the first genome-wide dataset on protein translation during fibroblast activation, which we were also able to follow-up in the diseased human heart," said Dr. Stuart Cook, a senior co-author of the study, the Tanoto Foundation Professor of Cardiovascular Medicine and Director of Duke-NUS' CVMD Programme, and a Senior Consultant at the National Heart Centre Singapore. "The combination of in-house experiments and large public data repositories, together with the novel functional genomics approach we used to integrate various data types, provide a powerful tool to explore gene regulation."
Professor Patrick Casey, Senior Vice Dean for Research at Duke-NUS, commented, "Our ability to understand disease is being revolutionized by the availability of new technologies, whose power can best be realized by interdisciplinary teams combined with the development of methods that can address previously intractable questions. As exemplified in this study, the best way to do this is by bringing together scientific and clinical expertise with cutting-edge technology."
The researchers recommend future research to carefully reveal the interdependencies and cross-talks in the various stages of gene expression in order to be able to develop a holistic view of the regulatory process contributing to the manifestation of disease.
More information: Sonia Chothani et al. Widespread Translational Control of Fibrosis in the Human Heart by RNA-Binding Proteins, Circulation (2019). DOI: 10.1161/CIRCULATIONAHA.119.039596