October 21, 2019 feature
Microbiome: Untapped source of novel antimicrobials

Just as Gold Rush prospectors once mined the Northern California hills for the shiny precious metal, "bioprospectors" are searching for a new prize: potential antimicrobial molecules—and they are hunting them down in the human microbiome.
For nearly two decades scientists have been lifting the veil of mystery from the microbial communities that call humans home. Now, they see the microbiome as an untapped source of molecules that may aid the fight against infections and possibly treat other disease processes.
A team of Princeton University bioprospectors led by Dr. Yuki Sugimoto has combined bioinformatics with synthetic biology to identify biologically active small molecules with drug-like capabilities. By using their new computational algorithm, Sugimoto and colleagues were able to illuminate the DNA fingerprints of potentially potent antimicrobial molecules.
"The human microbiome harbors thousands of bacterial species varied in composition between different sites of the human body, and between individuals," noted Sugimoto, a postdoctoral fellow in the laboratory of Dr. Mohamed Donia, a Princeton molecular biologist.
The microbiome is composed of a multitude of commensal microbes that reside in and on us, defined by vast colonies of bacteria, and even beneficial viruses and fungi that thrive symbiotically as part of each individual's personal ecosystem. Although remarkable progress has been made in recent years elucidating the effects of the microbiome on human health and disease, the underlying molecules and mechanisms governing these effects have remained largely unexplored—at least until now.
Princeton's bioprospecting research is a novel area in the growing arena of human microbiome investigations, which in recent years has moved from an arcane research focus to a multi-billion-dollar enterprise as biotechnology and pharmaceutical companies have begun projects.
Academically, the Human Microbiome Project, an initiative of the U.S. National Institutes of Health, which ran from 2007 to 2016, was an early effort designed to develop a "big picture" understanding of the role played by human microbial flora. A second phase, the Integrative Human Microbiome Project, which launched in 2014, is aimed at further characterizing commensal communities and their impact on disease processes, as well as their role in physical and mental health.
Sugimoto, whose research background is in synthetic biology, and who received a doctorate from Friedrich-Schiller University in Germany, has focused on identifying biologially active, microbiome-derived molecules. His latest data, reported in Science, sheds new light on the potency of the human microbiota and the potential role these microbes play in the development of antimicrobial medications of the future.
The research is not far-fetched, given the long and storied past of antibiotic development. For example, the antimicrobial activity of fungi has been known for nearly 100 years following Alexander Fleming's accidental discovery of penicillin in 1928. The 1943 Rutgers University discovery of the antibiotic streptomycin, a medication derived from the soil bacterium, Streptomyces griseus, and other findings, further laid the groundwork for studies demonstrating how seemingly simple organisms can produce complex, biologically active molecules. It has taken until now to grasp the potential of molecules that possess similar capability among the microbial communities of the microbiome.
Sugimoto and colleagues describe the human microbiome as an unparalleled resource for identifying the chemical structures of new drugs. "We discovered that members of a clinically used class of molecules are widely encoded in the human microbiome, and that they exert potent antibacterial activities against neighboring microbes," Sugimoto and the team wrote in Science. "Our approach paves the way toward a systematic unveiling of the chemical repertoire encoded by the human microbiome."
In their search, the investigators relied on their customized approach, a computer algorithm that allowed them to search a vast number of molecular fragments. This bioinformatics approach, MetaBGC (which stands for metagenome biosynthetic gene cluster), allowed them to uncover molecules never before reported from the human microbiome. They focused on samples from the mouth, gut and skin, biological sites rich in microbial colonies.
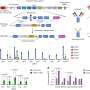
They found multiple novel enzymes, Type II polyketide synthase BGCs, or simply TII-PKS BGCs, from the three sites. TII-PKS molecules are well known because a TII-PKS enzyme is an important component in the drug doxorubicin, a cancer drug with antibiotic activity. Aside from its role in cancer treatment, doxorubicin is also used as an assay for screening environmental bacteria.
To test the importance of their finding about TII-PKS BGC molecules, Sugimoto and his team of bioprospectors tested samples from people worldwide: the United States, Denmark, Spain, Fiji and China. They found that regardless of geographic location, people possessed genes for these molecules, suggesting they are widespread in human populations.
"The structural diversity observed in the products of two of the 13 BGCs discovered here, and the fact that they resemble in structure or biological activity clinically used drugs, clearly motivates further functional investigations into this class," says Sugimoto.
"These investigations will not only serve as an important avenue for elucidating microbiome-host interactions at the molecular level, but they will also serve as an unprecedented resource for drug discovery from within the human body."
More information: Yuki Sugimoto et al. A metagenomic strategy for harnessing the chemical repertoire of the human microbiome, Science (2019) DOI: 10.1126/science.aax9176
© 2019 Science X Network