Transient and long-term disruption of gut microbes after antibiotics
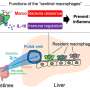
Trillions of microbes in the intestine aid human health, including digestion of breast milk, breaking down fiber and helping control the immune system. However, antibiotic treatment is known to disrupt the community structure of these microbes—500 to 1,000 bacterial species that have a mainly beneficial influence.
A study at the University of Alabama at Birmingham now has tracked this disruption at the level of a strain of microbes replacing another strain of the same species in 30 individuals—all of them young, healthy adults who would be expected to have stable microbial communities.
"As shown from our analysis, the capacity to recover with respect to the number and stability of new strains is specific for each individual," said Casey Morrow, Ph.D., leader of the research team and professor emeritus in UAB's Department of Cell, Developmental and Integrative Biology.
Differences in recovery have potential health implications, Morrow says.
"It is possible that as individuals age, with each differing in numbers and cycles of antibiotic treatment, the reservoir of microbial strains is depleted, resulting in an intra-individual recovery pattern for specific microbial strains," he said. "Understanding this recovery pattern—including the occurrence of particular strains following antibiotics—may be an important consideration for long-term health."
"In the future, the characterization of these individual-specific recovery patterns could also be used to forecast the susceptibility to both endogenous and exogenous microbial pathogens."
The UAB study used bioinformatic tools to analyze a previously described study of 18 individuals who had been given a single antibiotic, cefprozil, for a week. Their fecal samples were collected at pretreatment, at the end of antibiotic treatment and at three months post-treatment. The UAB study also analyzed previously described data of 12 individuals who were given a combination of three antibiotics—meropenem, gentamicin and vancomycin—for four days. Their fecal samples were collected at pretreatment; at end of treatment; and at four, 38 and 176 days post-treatment. Six control individuals who did not receive antibiotics were also analyzed.
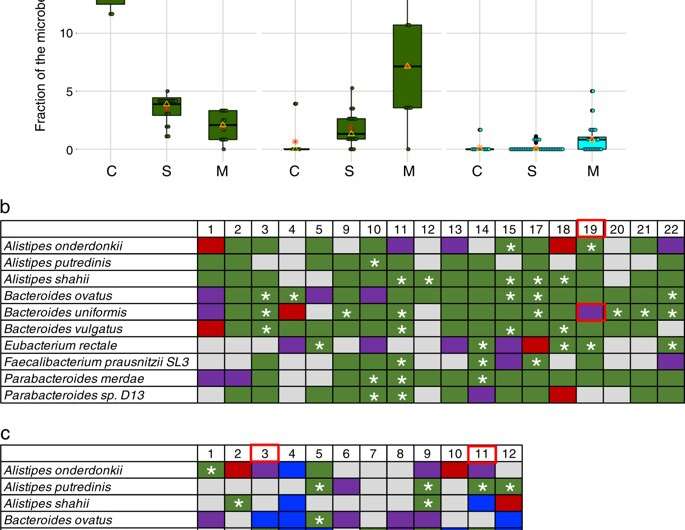
In general, the UAB researchers found that strains of the 10 most abundant species remained stable in controls. In the single antibiotic treatment individuals, 15 of 18 individuals had transient new strains post-treatment that, in turn, were replaced by the original strain by three months post-treatment.
In contrast, the triple-antibiotics individuals showed a significant increase of new strains that persisted as long as six months after treatment, as compared to the single antibiotic and the control individuals. Furthermore, the fraction of transient strains was also significantly higher in the multiple antibiotics individuals. This suggested a long-term change to an alternative stable microbiome state, Morrow says. These changes were not due to a difference in growth rates.
"Given the importance of the microbiome in human health, we think our results with these data sets can be used to help evaluate microbiome stability under different conditions," Morrow said. "For example, we can now provide guidance to clinical investigators to judge the impact of certain treatments for diseases, such as cancer or diabetes, on the gut microbial community that could be significant for evaluation of outcomes. Furthermore, this approach could be applied to a patient's pre- and post-hospitalization to identify individuals who may need further management of their microbiomes."
"This study used a strain-tracking bioinformatics tool previously developed by UAB, called Window-based Similarity Single-nucleotide-variant, or WSS, for tracking of individuals' microbial strains from pre-treatment to post-antibiotics treatment," said Hyunmin Koo, Ph.D., UAB Department of Genetics and Heflin Center for Genomic Science, who led the informatics analysis. "This technique advances analysis of the impact of antibiotics on the human gut microbiota. Previous studies of the microbiome had been able to determine an overall taxonomic profile including the relative abundance information of each species, but showed a limitation to distinguish each species at the strain level or track the same strain in each individual at the longitudinal level.
In 2017, the UAB researchers used WSS to show the first direct demonstration that fecal donor microbes—used to treat patients with recurrent Clostridium difficile infections—remained in recipients for months or years after fecal transplants.
The antibiotics study was published in npj Biofilms and Microbiomes.
More information: Hyunmin Koo et al. Individualized recovery of gut microbial strains post antibiotics, npj Biofilms and Microbiomes (2019). DOI: 10.1038/s41522-019-0103-8