How HIV develops resistance to key drugs discovered
The mechanism behind how HIV can develop resistance to a widely-prescribed group of drugs has been uncovered by new research from the Crick and Dana-Farber Cancer Institute, with the findings opening the door to the development of more effective treatments.
Today, a number of drugs are available which help to control HIV infection, including a group called integrase strand transfer inhibitors. There are four drugs within this family of medication: raltegravir, elvitegravir, dolutegravir and bictegravir. They all work by binding with one of HIV's key enzymes, called integrase, to stop it from inserting the virus' genetic information into DNA of human cells. While initially highly effective, over time HIV can develop resistance to these drugs.
The study, published on-line in Science 30 January 2020, discovered the mechanism HIV uses to develop resistance to this group of drugs. Although the drugs are normally very effective at binding and blocking integrase, over time, the virus can weaken this bond and thus enable its key enzyme to work again.
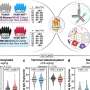
The researchers at the Francis Crick Institute in London uncovered this by exploring the structure of integrase from a virus that is highly similar to the ancestor of HIV, using cryo-electron microscopy. This technique uses a powerful microscope which fires electrons at a frozen sample of the drug-enzyme complex. By recording how the electrons interact with the samples, the researchers created detailed images, at a nearly atom-by-atom level.
"The unusual property of these drugs is that they interact with metal ions, which normally allows them to make very strong bonds to the viral enzyme's active site. We found that HIV can subtly alter the chemical environment of the metals, and as if using a remote control, reduce the strength of drug binding. This is an unexpected chink in the armour of strand transfer inhibitors," says Peter Cherepanov, co-lead author and group leader in the Chromatin and Mobile DNA Laboratory at the Crick and Professor of Molecular Virology at Imperial College London.
"The good news is that we have finally visualised the precise structure of the viral enzyme's active site, right where the drugs bind. These blueprints will inform the design of more effective integrase inhibitors that could improve the lives of the many millions of people living with HIV," says Alan Engelman, co-lead author from the Department of Cancer Immunology and Virology at the Dana-Farber Cancer Institute and Professor of Medicine at Harvard Medical School in Boston, USA.
"The weakening of drug binding occurs due to a combined effect of mutations and a loss of key water molecules in the active site. Understanding this mechanism will help improve this class of drugs in the future," comments Edina Rosta, co-author from the Crick and Reader of Computational Chemistry at Kings College London, whose team conducted complex computations on the integrase structures.
Daniel Kuritzkes, Chief of Infectious Diseases at Brigham and Women's Hospital and Professor of Medicine at Harvard Medical School, who was not directly involved in the study, adds: "The work by the Cherepanov and Engelman laboratories importantly informs the mechanism of resistance to a class of drugs that are now recommended first-line treatment for HIV worldwide."
"This research is an outstanding example of how we can use cryo-electron microscopy to reveal the intricate relationships between drugs and their targets, providing results that could lead to clinical benefit," explains Peter Rosenthal, the head of the Structural Biology of Cells and Viruses laboratory at the Francis Crick Institute.
More information: "Structural basis of second-generation HIV integrase inhibitor action and viral resistance" Science (2020). science.sciencemag.org/lookup/ … 1126/science.aay4919