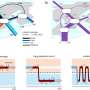
Release of 216 SARS-CoV-2 genomes provides insights into transmission cluster in Austria
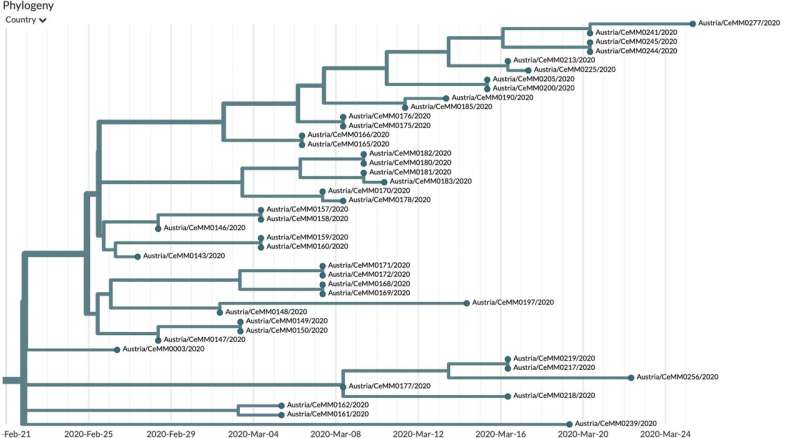
Two-hundred-sixteen SARS-CoV-2 genome sequences have now been completed and released in the framework of the Mutational Dynamics of SARS-CoV-2 in Austria project from CeMM, the Research Center for Molecular Medicine of the Austrian Academy of Sciences, in close collaboration with the Medical University of Vienna, the Medical University of Innsbruck and the Austrian Agency for Health and Food Safety (AGES). This data represents another milestone in the project and will help to understand the mutational landscape and evolution of the Austrian SARS-CoV-2 strains. Together with this release, CeMM researchers have published a dedicated website that offers background information as well as interactive access to the data for scientists and laypeople alike. The virus genomes can be explored interactively and intuitively via advanced visualization tools provided by CeMM and the open source project Nextstrain.
The spread of the pandemic SARS-CoV-2 virus in the human population is met with unprecedented collaborative scientific efforts around the world to unravel its virological, immunological and disease-causing properties. Genome sequences of SARS-CoV-2 viruses circulating worldwide are being published and made openly available to the international scientific community. A better understanding of the viral evolution will be instrumental to comprehend the underlying mutational dynamics and support the development of effective antiviral treatment and vaccine strategies to halt the COVID-19 pandemic.
In Austria, CeMM researchers with collaboration partners from the Medical University of Vienna were pioneers in sequencing the first genomes obtained from Austrian patient-derived samples. Within the scope of the "Mutational Dynamics of SARS-CoV-2 in Austria" project, the first samples were made available to the public and the international research community already at the beginning of April 2020. This highly collaborative initiative, led by CeMM Principal Investigators Andreas Bergthaler and Christoph Bock, aims at elucidating 1,000 SARS-CoV-2 genomes in Austrian patients using cutting-edge next-generation sequencing techniques and sophisticated computational analyses. The project includes a wide national collaboration network with partners from the Medical University of Vienna (Judith Aberle, Stephan Aberle, Elisabeth Puchhammer-Stöckl), the Medical University of Innsbruck (Wegene Borena, Dorothee von Laer; Manfred Nairz, Günter Weiss) and the Austrian Agency for Health and Food Safety (Daniela Schmid, Peter Hufnagl) as well as several hospitals and diagnostic laboratories.
-
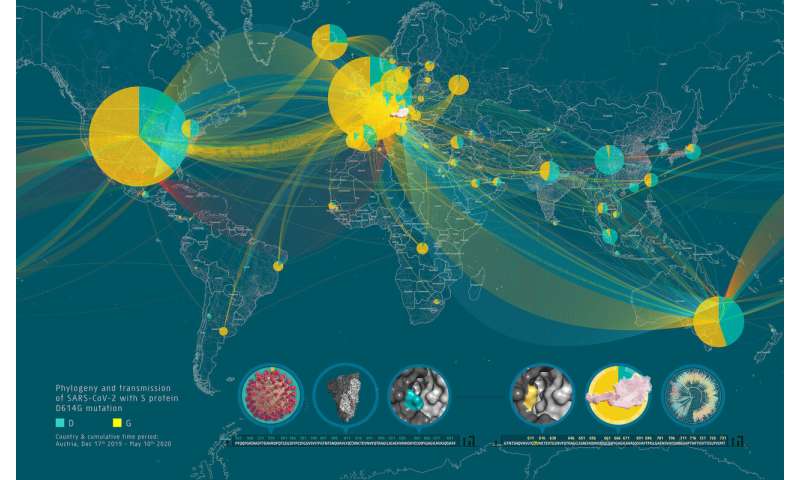
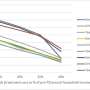
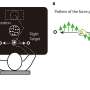
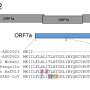
The D614G mutation in the viral S protein represents an early branching point that is dominant in the European infection clusters and parts of North America. Sequence analysis suggests further transmission of this strain to North America and subsequent reintroduction events from the USA to many European countries. Most of the Austrian patient samples show the same S D614G mutation shared by many other European strains. The Illustration 2 was generated by data-driven vector-graphics extracted from Nextstrain.org, enabled by data from GISAID.org, world map-data from Openstreetmap.org and 3D elements rendered with Octane renderer for Unity3D. Credit: Bobby Rajesh Malhotra / CeMM (Creative Commons license: CC BY-NC) -
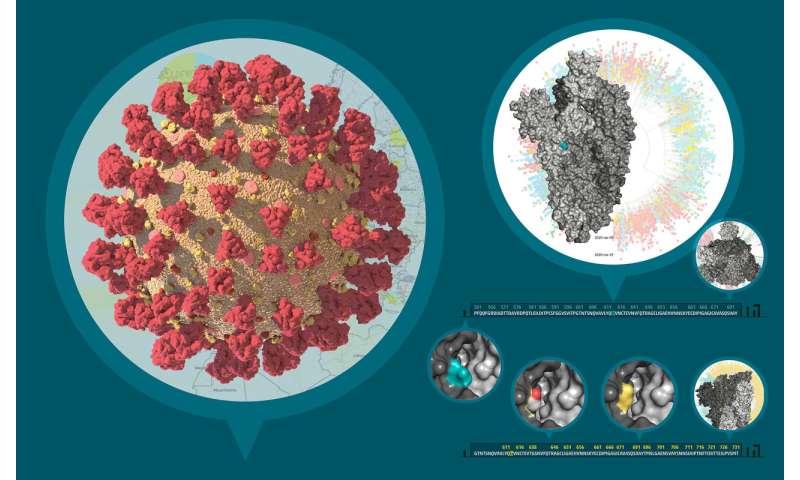
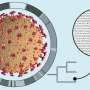
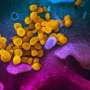
To the left, this composite image depicts the three-dimensional structure of SARS-CoV-2 virion based on custom mesoscale 3D-model of SARS-CoV-2 with RCSB PDB 3D-surface data and experimental protein structures (PDB: 6VSB, 6VXX, 6VXY). To the top right, the three-dimensional structure of the viral S protein and a phylogenetic tree of international SARS-CoV-2 genomes is shown. To the bottom right, the predicted conformational change in the S protein structure caused by the D614G mutation is shown.The Illustration 3 was generated by data-driven vector-graphics extracted from Nextstrain.org, enabled by data from GISAID.org, world map-data from Openstreetmap.org and 3D elements rendered with Octane renderer for Unity3D. Credit: Bobby Rajesh Malhotra / CeMM (Creative Commons license: CC BY-NC)
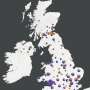
Building on their first 21 genome sequences published on 3 April 2020, this second release with 216 new SARS-CoV-2 genomes from across Austria provides a more detailed picture about the circulating viruses and the early phase of the COVID-19 pandemic. The genetic differences of the genomes indicate that the pool of circulating viruses in the early phase of the pandemic was already highly diverse, with some viruses leading to bigger transmission clusters than others. Importantly, Austria has benefited from painstaking contact tracing by the Austrian Agency for Health and Food Safety, which defined more than 250 clusters of COVID-19 cases.
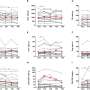
"Our integrative analysis of the new SARS-CoV-2 genome information with epidemiological data provides valuable new insights into how the virus has been spreading through the country. The virus sequence data provides independent support for many of the epidemiological findings resulting from contact tracing. At the same time, we observed both heterogeneity within clusters and obtained evidence for multiple viruses circulating at the same time," says Andreas Bergthaler, project co-coordinator and CeMM Principal Investigator (see illustration 1).
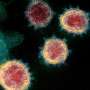
The molecular sequence analysis revealed on average 6.9 mutations per viral genome, of which more than 4 resulted in changed amino acids. A particular focus of interest worldwide rests on mutations in the viral S or spike protein, which decorates the surface of the virion and lends the coronavirus its eponymous crown-like appearance (see illustration 2). This viral S protein is essential for binding to the cellular receptor ACE2 as well as the presumed primary target for neutralizing antibodies. As such, mutations in this region will be important for the development of serological tests and antibody-mediated protection by vaccines. Within all Austrian SARS-CoV-2 genomes, we identified a total number of 12 mutated residues in the 1,273 amino acid long S protein. One particular mutation in the S protein, D614G, represents an early branching point that is dominant in the European infection clusters and parts of North America (see illustrations 2 and 3). This mutation is also found in many of the Austrian samples. The potential functional consequences of the D614G mutation are currently being investigated by several international laboratories. CeMM's ongoing in-depth viral mutation analyses with an interdisciplinary team from the fields of genomics, biomathematical modeling, evolution biology and virology will yield new insights into how the S protein and other viral proteins evolve as well as how these mutations are transmitted between individuals.
Today's release marks yet another crucial milestone as CeMM intensifies its efforts to communicate science and to reach out to the general public. Based on the tools of the open source project Nextstrain.org, CeMM researchers released a Nextstrain Austria version that allows users to compare and visualize all sequenced Austrian virus genomes with more than 8,000 virus genomes from all over the world. The Nextstrain Austria Blog will provide concise up-to-date and data-driven stories to share interesting insights about what the virus genomes tell us about the SARS-CoV-2 pandemic. These narratives will explore the actual genome data generated by CeMM and its partners, and allow users to explore the content interactively with an intuitive design. Users are guided through different dashboards where they encounter interactive graphics with a short explanatory text on different topics related to the SARS-CoV-2 virus. The visualization tool is available in both English and German and can be accessed at https://cemm.at/SARS-cov-2/. The blog will be continuously updated and give first-hand insights into ongoing intense research about the SARS-CoV-2 virus.
The project team will continue their efforts to sequence 1,000 SARS-CoV-2 viral genomes, which will provide a detailed picture of emerging viral mutations and virus transmissions. This will offer further valuable scientific insights to epidemiologists, health care professionals and public health experts to assess transmission routes of the virus as well as its potential to subvert vaccine-induced immune responses and to acquire resistance against antiviral drugs.