May 28, 2020 report
Researchers map structural variants in 17,795 sequenced human genomes
A large team of researchers affiliated with a very large number of medical institutions across the U.S. and also three from Finland, two from Mexico, and one from the U.K. has mapped structural variants in 17,795 sequenced genomes. In their paper published in the journal Nature, the group describes their work, which was meant to explore structural variants in genome sequencing, and what they found.
As the researchers note, one of the main objectives when scientists conduct whole-genome sequencing on a large number of people is to learn more about genetic variations. Such variations refer to alterations that may be pathogenic, benign or of unknown significance. Genome variants are classified as small insertion-deletion, single nucleotide or structural. They further note that research into structural variants has lagged behind that of the others, mostly due to a lack of tools and resources dedicated to their study.
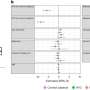
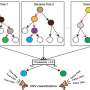
In this new effort, they sought to improve on such work by using what they describe as a scalable pipeline to map and characterize structural variants in 17,795 deeply sequenced genomes. The scalable pipeline refers to a software toolkit they developed that allows for workflow and large-scale structural variant callset generation—it produced data describing resolution-aware cross-sample merging, per-sample variant discovery, variant classification, breakpoint genotyping and copy number annotation.
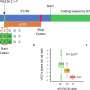
The researchers used the toolkit to analyze the genomes of 17,795 people of European, Latino and African ancestry—the genome data was obtained from several publicly accessible databases—and each genome was sequenced to a minimum 20-fold coverage. They note that the size of the sample and the use of deep sequencing allowed them to map rare variants with high resolution. It also allowed them to generate estimates of the relative burden of deleterious structural variants. On average, they saw deletions most often in the genomes, followed by insertions. There were 2.9 rare structural variants on average (in protein-coding portions of the genome) per person in the dataset. The toolkit estimated that 17 percent of the rare variants that led to loss of function could be blamed on structural variants.
More information: Mapping and characterization of structural variation in 17,795 human genomes, Nature (2020). DOI: 10.1038/s41586-020-2371-0
© 2020 Science X Network