New way to analyze fMRI data offers path to improving treatment for schizophrenia
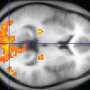
Researchers at the University of Maryland, Baltimore County (UMBC) have developed tools to improve the analysis of functional magnetic resonance imaging (fMRI) data. Tülay Adali, professor of computer science and electrical engineering and director of UMBC's Machine Learning for Signal Processing Lab, and Qunfang Long, a Ph.D. candidate at UMBC in electrical engineering, have spearheaded groundbreaking work identifying key patterns in brain imaging for those with particular mental illnesses, such as schizophrenia. This new research is published in NeuroImage Volume 216. Their work can assist in diagnosis and treatment of patients with mental illnesses that can be difficult to identify. It can also show medical practitioners whether the current treatments have or have not been working based on image groupings.
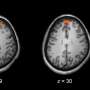
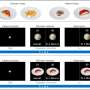
The image analysis method developed by Adali and Long is called independent vector analysis (IVA) for common subspace extraction (CS). Through this method, they were able to categorize subgroups of fMRI data based solely on brain activity, proving that there is a connection between brain activity and certain mental illnesses. In particular, they were able to identify subgroups of schizophrenia patients using the fMRI data that they analyzed.
Previously, there was not a clear way to group schizophrenia in patients based on brain imaging alone, but the methods developed by Adali and Long show that there is a significant connection between a patient's brain activity and their diagnoses. "The most exciting part is that we found out the identified subgroups possess clinical significance by looking at their diagnostic symptoms," explains Long. "This finding encouraged us to put more effort into the study of subtypes of patients with schizophrenia using neuroimaging data."
Importantly, the IVA-CS method used to identify these subgroups also preserves nuances in the data, but still renders statistically significant groupings. "Now that data-driven methods have gained popularity, a big challenge has been capturing the variability for each subject while simultaneously performing analysis on fMRI datasets from a large number of subjects. Now we can perform this analysis effectively, and can identify meaningful groupings of subjects," says Adali.
Diagnosing and treating mental illness is incredibly complex. The same illness will present differently in different patients, and there is often no singular treatment that will be effective for all patients. Once a treatment is in place, determining if it is successful can also vary by patient. Adali and Long's research along with their long-time collaborator Vince Calhoun at the Tri-institutional Center for Translational Research in Neuroimaging and Data Science in Georgia, U.S. responds to this variability by giving medical practitioners an objective way to analyze the fMRI results for patients within relatively homogenous diagnostic subgroups, and then compare fMRI results over time for the same patient. Consider a schizophrenic patient who receives treatment and returns in six months to be evaluated again. If their fMRI data resembles that of the control group of mentally healthy patients more than that of other patients with schizophrenia, that is objective evidence that the treatment is working. On a larger scale, this data provides a better look at patients' medical outcomes as a result of treatment.
To further improve upon this groundbreaking analysis of fMRI data, Adali's team will work with longitudinal data to determine which treatments work best for subgroups of patients with specific mental illnesses. This method will also be used in a longitudinal study of adolescents to see whether there are links between fMRI images and the addiction and substance use patterns of those adolescents over time.
More information: Qunfang Long et al, Independent vector analysis for common subspace analysis: Application to multi-subject fMRI data yields meaningful subgroups of schizophrenia, NeuroImage (2020). DOI: 10.1016/j.neuroimage.2020.116872