St. Jude Cloud portal expands access to treasure trove of pediatric solid tumor data
An innovative, interactive cloud-based data portal debuted this week that lets academic researchers mine the world's largest and most comprehensive collection of scientific resources for studying pediatric solid tumors and their related biology. The Childhood Solid Tumor Network (CSTN) data portal on St. Jude Cloud was created to improve access to the detailed data available through the network, stimulating the research and development of novel, lifesaving therapies.
The CSTN includes wide-ranging data on 170 patient-derived samples representing 21 different childhood solid tumors, including neuroblastoma, rhabdomyosarcoma and rare tumors. The samples are orthotopic patient-derived xenografts, meaning the human tumor samples are implanted and grown in the corresponding location in mice.
Tumor samples and related data are shared with academic researchers at no charge and with no obligation to collaborate. Michael Dyer, Ph.D., chair of the St. Jude Developmental Neurobiology department, and Alberto Pappo, M.D., of the St. Jude Oncology department, launched the network in 2013. The goal was to accelerate discoveries in the field. "There had been no significant improvement in outcomes for children with solid tumors in the past three decades," said Dyer, who is a Howard Hughes Medical Institute investigator. Pediatric solid tumors are developmental tumors. To understand how the tumor begins, grows and spreads, they need to be studied in the environment in which they develop.
"As interest in the network grew, we realized scientists needed to dig deeper into the data," Dyer said. "We created the interactive browser to make that possible." The CSTN data portal was unveiled this week at the annual meeting of the American Association for Cancer Research, which was held virtually.
Web-based data
The portal makes it easier for researchers to download and analyze the data themselves.
Available data and analytic tools include:
- Next-generation, whole genome sequencing of the patient tumors, germline and the orthotopic patient-derived xenografts. These are presented as an interactive application, allowing for customized heatmaps to represent mutations and individual gene searches. Raw genomic sequences are also uploaded and available.
- Interactive browsers for epigenetic and, in some cases, proteomic data
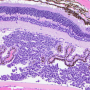
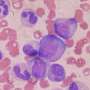
- Histology images with tissue-specific stains and electron microscopy images at three magnifications for each sample
- Clonal analysis of the matched patient samples and orthotopic xenografts
- Sample-level search and visualization of specific characterization data that let users filter by factors such as patient's age at diagnosis or primary versus recurrent tumors
Preclinical pharmacokinetic reports, tumor propagation protocols and more
Single-cell RNA sequencing of patient and patient derived-orthotopic xenografts is underway. Additional tumor samples and data are added to the network as they become available.
Academic researchers can also request cryopreserved cells from orthotopic patient-derived xenografts as well as fresh frozen tissue or cells and formalin-fixed paraffin-embedded tissue blocks.
As of June 2020, more than 200 researchers from more than 100 institutions worldwide have requested and received more than 1,500 tumor samples from the network.
"We feel the network and data portal are the best way to make the most of the tumor tissue samples that children and their parents have donated to provide resources to speed cures and understand the diseases," said Asa Karlstrom, Ph.D., of Developmental Neurobiology. She was co-first author of a report on the network data portal that was included in an AACR poster session.