New cause of syndromic microcephaly identified

Greenwood Genetic Center (GGC) researchers, in collaboration with scientists from Belgium, Spain, and Italy, have confirmed that variants in LMNB1 cause syndromic microcephaly with variable short stature, intellectual disability, and other neurological symptoms. The findings are reported in the October issue of the American Journal of Human Genetics.
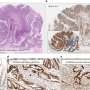
The paper describes seven patients from five families with novel de novo variants in LMNB1, all with pronounced primary microcephaly as the core feature. Variants were identified through whole exome sequencing.
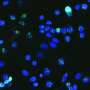
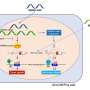
Functional analysis of each of the gene variants and resultant lamin B1 protein showed that these variants act in a dominant fashion to disrupt nuclear envelope integrity which in turn causes the nucleus to become misshapen.
"We used a HeLa cell system to analyze the three missense mutations in this gene and demonstrated impaired function of the nuclear lamina," said Tonya Moss, GGC research technologist and co-lead author along with Francesca Cristofoli, Ph.D., of University Hospitals Leuven in Belgium. "Two other variants displayed misshapen nuclei and a decrease in nuclear localization of the lamin B1 protein, and another showed an increased frequency of condensed nuclei."
Duplications in LMNB1 are a known cause of autosomal dominant adult-onset leukodystrophy, but this paper is the first to show that variants in this gene can cause congenital abnormalities through nuclear envelope disruption.
"Because this gene product has multiple domains, it is likely that different variants in LMNB1 will work by different mechanisms, necessitating a case-by-case analysis of pathogenicity, which is important for future treatment studies as well as reproductive planning for families," said Richard Steet, Ph.D., co-corresponding author and Director of Research at the JC Self Research Institute at GGC. "These findings add LMNB1 to the growing list of genes implicated in severe autosomal dominant microcephaly and broadens the phenotypic spectrum of the laminopathies."
More information: Francesca Cristofoli et al, De Novo Variants in LMNB1 Cause Pronounced Syndromic Microcephaly and Disruption of Nuclear Envelope Integrity, The American Journal of Human Genetics (2020). DOI: 10.1016/j.ajhg.2020.08.015