Researchers develop tool to better predict treatment course for lung cancer
Personalized treatment options for patients with lung cancer have come a long way in the past two decades. For patients with non-small cell lung cancer, the most common subtype of lung cancer and the leading cause of cancer-related death worldwide, two major treatment strategies have emerged: tyrosine kinase inhibitors and immune checkpoint inhibitors. However, choosing the right therapy for a non-small cell lung cancer patient isn't always an easy decision, as biomarkers can change during therapy rendering that treatment ineffective. Moffitt Cancer Center researchers are developing a noninvasive, accurate method to analyze a patient's tumor mutations and biomarkers to determine the best course of treatment.
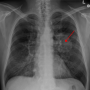
In a new article published in Nature Communications, the research team demonstrates how a deep learning model using positron emission tomography/computerized tomography radiomics can identify which non-small cell lung cancer patients may be sensitive to tyrosine kinase inhibitor treatment and those who would benefit from immune checkpoint inhibitor therapy. The model uses PET/CT imaging with the radiotracer 18F-Fluorodeoxyglucose, a type of sugar molecule. Imaging with 18F-FDG PET/CT can pinpoint sites of abnormal glucose metabolism and help accurately characterize tumors.
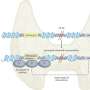
"This type of imaging, 18F-FDG PET/CT, is widely used in determining the staging of patients with non-small cell lung cancer. The glucose radiotracer used is also known to be affected by EGFR activation and inflammation," said Matthew Schabath, Ph.D., associate member of the Cancer Epidemiology Department. "EGFR, or epidermal growth factor receptor, is a common mutation found in non-small cell lung cancer patients. EGFR mutation status can be a predictor for treatment, as patients with an active EGFR mutation have better response to tyrosine kinase inhibitor treatment."
For the study, the Moffitt team developed an 18F-FDG PET/CT-based deep learning model using retrospective data from non-small cell lung cancer patients at two institutions in China: Shanghai Pulmonary Hospital and Fourth Hospital of Hebei Medical University. The model classifies EGFR mutation status by generating an EGFR deep learning score for each patient. Once created, the researchers further validated the model using patient data from two additional institutions: Fourth Hospital of Harbin Medical University and Moffitt Cancer Center.
"Prior studies have utilized radiomics as a noninvasive approach to predict EGFR mutation," said Wei Mu, Ph.D., study first author and postdoctoral fellow in the Cancer Physiology Department. "However, compared to other studies, our analysis yielded among the highest accuracy to predict EGFR and had many advantages, including training, validating and testing the deep learning score with multiple cohorts from four institutions, which increased its generalizability."
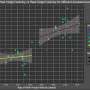
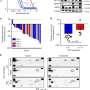
"We found that the EGFR deep learning score was positively associated with longer progression free survival in patients treated with tyrosine kinase inhibitors, and negatively associated with durable clinical benefit and longer progression free survival in patients being treated with immune checkpoint inhibitor immunotherapy," said Robert Gillies, Ph.D., chair of the Cancer Physiology Department. "We would like to perform further studies but believe this model could serve as a clinical decision support tool for different treatments."
More information: Wei Mu et al, Non-invasive decision support for NSCLC treatment using PET/CT radiomics, Nature Communications (2020). DOI: 10.1038/s41467-020-19116-x