This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
AI predicts tumor-killing cells with high accuracy, study shows
Using artificial intelligence, Ludwig Cancer Research scientists have developed a powerful predictive model for identifying the most potent cancer-killing immune cells for use in cancer immunotherapies.
Combined with additional algorithms, the predictive model, described in the current issue of the journal Nature Biotechnology, can be applied to personalized cancer treatments that tailor therapy to the unique cellular makeup of each patient's tumors.
"The implementation of artificial intelligence in cellular therapy is new and may be a game-changer, offering new clinical options to patients," said Ludwig Lausanne's Alexandre Harari, who led the study with graduate student Rémy Pétremand.
Cellular immunotherapy involves extracting immune cells from a patient's tumor, optionally engineering them to enhance their natural abilities to combat cancer and reintroducing them to the body after they've been expanded in culture. T cells are one of the two main types of white blood cells, or lymphocytes, that circulate in the blood and patrol for virally infected or cancerous cells.
T cells that penetrate solid tumors are known as tumor-infiltrating lymphocytes, or TILs. However, not all TILs are effective at recognizing and attacking tumor cells.
"Only a fraction is in fact tumor reactive—the majority are bystanders," Harari explained. "The challenge we set for ourselves was to identify the few TILs that are equipped with T cell receptors able to recognize antigens on the tumor."
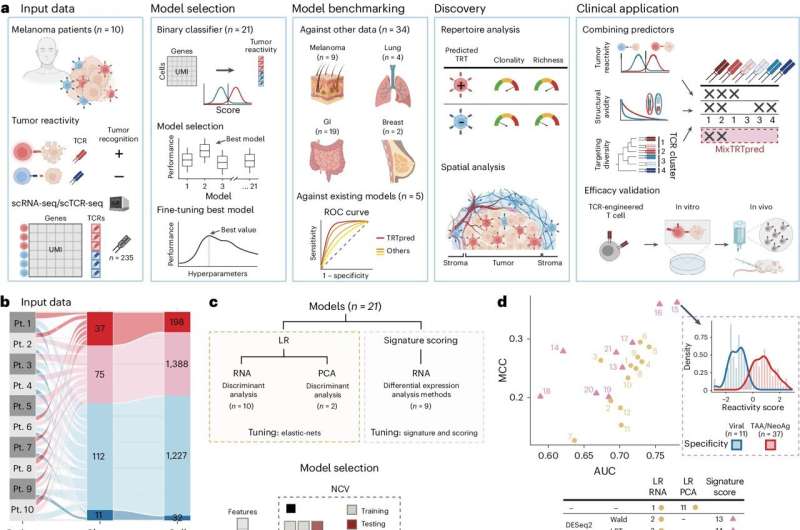
To do this, Harari and his team developed a new AI-driven predictive model, called TRTpred, that can rank T cell receptors (TCRs) based on their tumor reactivity. To develop TRTpred, they used 235 TCRs gathered from patients with metastatic melanoma, already classified as either tumor-reactive or non-reactive.
The team loaded the global gene-expression—or transcriptomic—profiles of the T cells carrying each TCR into a machine learning model to identify patterns that differentiate tumor-reactive T cells from inactive counterparts.
"TRTpred can learn from one T cell population and create a rule which can then be applied to a new population," Harari explained. "So, when faced with a new TCR, the model can read its transcriptomic profile and predict whether it is tumor reactive or not."
The TRTpred model analyzed TILs from 42 patients with melanoma and gastrointestinal, lung and breast cancer and identified tumor-reactive TCRs with about 90 percent accuracy. The researchers further refined their TIL selection process by applying a secondary algorithmic filter to screen for only those tumor-reactive T-cells with "high avidity"—that is, those that bind strongly to tumor antigens.
"TRTpred is exclusively a predictor of whether a TCR is tumor reactive or not," Harari explained. "But some tumor-reactive TCRs bind very strongly to tumor cells and are therefore very effective, while others only do so in a lazy way. Distinguishing the strong binders from the weak ones translates into efficacy."
The researchers demonstrated that T cells flagged by TRTpred and the secondary algorithm as both tumor-reactive and having high avidity were more often found embedded within tumors rather than in the adjacent supportive tissue, known as stroma. This finding aligns with other research showing that effective T cells typically penetrate deep into tumor islets.
The team then introduced a third filter to maximize recognition of diverse tumor antigens. "What we want is to maximize the chances the TILs will target as many different antigens as possible," Harari said.
This final filter organizes TCRs into groups based on similar physical and chemical characteristics. The researchers hypothesized that TCRs in each cluster recognize the same antigen. "So, we pick within each cluster one TCR to amplify, so that we maximize the chances of distinct antigen targets," said Vincent Zoete, a computational scientist at Ludwig Lausanne who developed the TCR avidity and the TCR clustering algorithms.
The researchers call the combination of TRTpred and the algorithmic filters MixTRTpred.
To validate their approach, Harari's team cultivated human tumors in mice, extracted TCRs from their TILs and used the MixTRTpred system to identify T cells that were tumor-reactive, had high avidity and targeted multiple tumor antigens. They then engineered T cells from the mice to express those TCRs and showed that these cells could eliminate tumors when transferred into the mice.
"This method promises to overcome some of the shortcomings of current TIL based therapy, especially for patients dealing with tumors not responding to such therapies today," said Ludwig Lausanne Director George Coukos, a co-author of the study who is planning to launch a Phase I clinical trial that will test the technology in patients.
"Our joint efforts will bring forth a completely new type of T cell therapy."
More information: Rémy Pétremand et al, Identification of clinically relevant T cell receptors for personalized T cell therapy using combinatorial algorithms, Nature Biotechnology (2024). DOI: 10.1038/s41587-024-02232-0