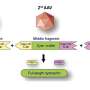
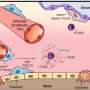
Germline whole exome sequencing reveals potential role of hereditary predisposition in small cell lung cancer
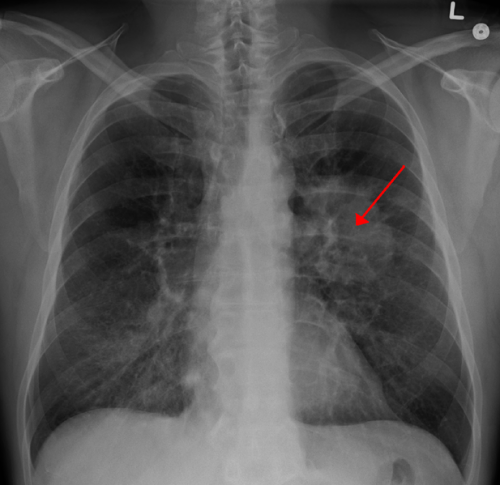
A study presented today by Dr. Nobuyuki Takahashi of the Center for Cancer Research (CCR), National Cancer Institute (NCI), Bethesda, Md. at the IASLC World Conference on Lung Cancer Singapore demonstrates that small cell lung cancer (SCLC) may have an inherited predisposition and lays the foundation for understanding the interaction between genotype and tobacco exposure in exacerbating SCLC risk as well as potential therapeutic implications. Because tobacco is the dominant carcinogen, secondary causes of lung cancer are often diminished in perceived importance, especially in SCLC, the most lethal lung cancer. SCLC is almost exclusively related to tobacco and comprises 15% to 20% of all lung cancers.
The study was conducted by researchers at the NCI CCR including Drs. Takahashi, Camille Tlemsani, and Lorinc Pongor, and led by Dr. Anish Thomas of the Developmental Therapeutics Branch. To explore the genetic basis of SCLC, they sequenced germline whole exomes of 87 patients (77 with SCLC, 10 with extrapulmonary small cell) and compared these with an independent SCLC cohort and with cancer-free non-Finnish European individuals from the Exome Aggregation Consortium (ExAC) cohort. They also evaluated clinical characteristics associated with the germline genotype.
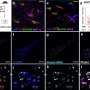
Among 607 cancer predisposition and SCLC-related genes, the researchers discovered 42 deleterious variants in 35 genes among 38 (43.7%) of patients. Identification of variants influenced medical management and family member testing in 9 (10.3%) patients. Six germline mutations were also found in the independent cohort of 79 patients with SCLC, including three of the same variants (MUTYH G386D, POLQ I421Rfs*7, and RNASEL E265X). By tumor whole exome sequencing they confirmed loss of heterozygosity of MLH1, BRCA2, and SMARCA4 genes.
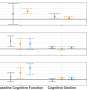
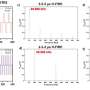
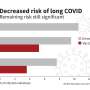
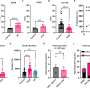
Consistent with the contribution to potential cancer predisposition, patients with MLH1, BRCA2, and MUTYH germline mutations had multiple personal and family history of cancer and lung cancers including SCLC. Unselected patients with SCLC in the cohort were more likely to carry germline RAD51D, CHEK1, BRCA2, and MUTYH mutations than healthy controls in the ExAC cohort. Pathogenic germline mutations were significantly associated with the likelihood of first-degree relatives with cancer or lung cancer (odds ratio: 1.82, 2.60, respectively) and longer recurrence-free survival following platinum-based chemotherapy (hazard ratio: 0.46, p = 0.002), independent of known prognostic factors including sex, stage, and age at diagnosis. Finally, they tested therapeutic relevance of these observations in a SCLC patient with pathogenic BRIP1 mutation, who achieved a tumor response (-64% decrease in tumor size) with a combination treatment of a topoisomerase 1 inhibitor and a poly (ADP-ribose) polymerase inhibitor.
"The study opens new avenues for directed cancer screening for patients and their families, as well as subtyping and targeted therapies of SCLC, currently treated as a single entity," Dr. Takahashi reported.
The study is published in Science Translational Medicine.
More information: "Whole-exome sequencing reveals germline-mutated small cell lung cancer subtype with favorable response to DNA repair–targeted therapies" Science Translational Medicine (2021). stm.sciencemag.org/lookup/doi/ … scitranslmed.abc7488