Coloring tumors reveals their bad influence
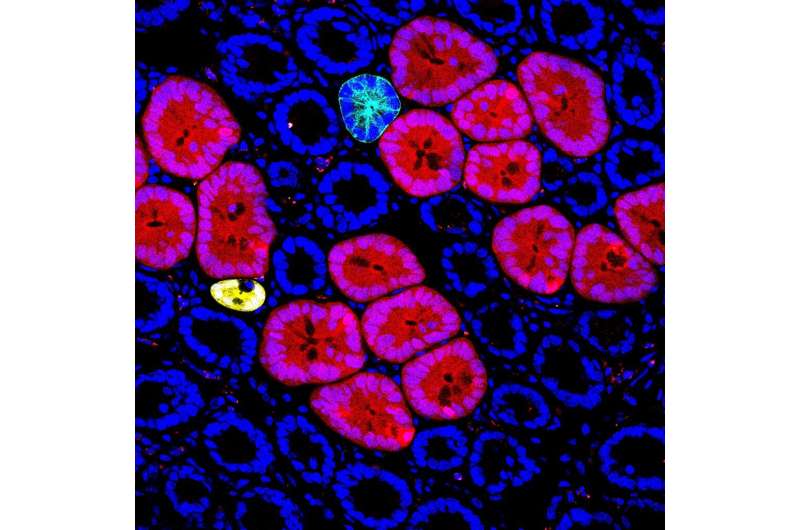
Red2Onco, an innovative genetic mouse model, allows to detect the very initial steps that lead to cancer development. Red2Onco's multicolor labeling system allows to trace intestinal tumor development after the first oncogenic hit at the single cell level. The research carried out at IMBA—Institute of Molecular Biotechnology of the Austrian Academy of Sciences—and the University of Cambridge is now published in the journal Nature.
Studies on cancer are limited by the threshold at which cellular transformations become clinically detectable. However, the very initial phase on the way to malignancy is histologically invisible, as the process originates from one single cell. In this early phase, a so-called "seeding cell" acquires an initial pro-cancerous mutation, also known as the "first oncogenic hit," while being completely surrounded by normal tissue. To overcome the detection barrier, a team of researchers around IMBA group leader Bon-Kyoung Koo and University of Cambridge group leader Professor Benjamin D. Simons developed a laboratory system to dissect the pre-cancerous steps that remained under the radar up until present.
On the hunt for the roots of cancer
"With the advance of technologies like deep targeted DNA sequencing, researchers noticed that cancer-associated mutations are already present in normal tissues, which is very scary," states Bon-Kyoung Koo, referring to the turning point that set him, like a forensic detective, on the traces of the first oncogenic hit.
As most human cancer types originate in the epithelium, Koo, a geneticist, decided to build a genetic model to examine the effect of the first oncogenic transformations in the mouse gut. This genetic model, which the team calls Red2Onco, is a multi-color labeling technology that allows to track the initiation of tumorigenesis from a single mutant cell. With Red2Onco, the team found that mutant cells create a hostile environment to their neighboring non-mutant cells and massively deregulate the normal stem cell niche in the mouse intestinal tissue. "This is by far the most exciting approach that we have tried," says Bon-Kyoung Koo.
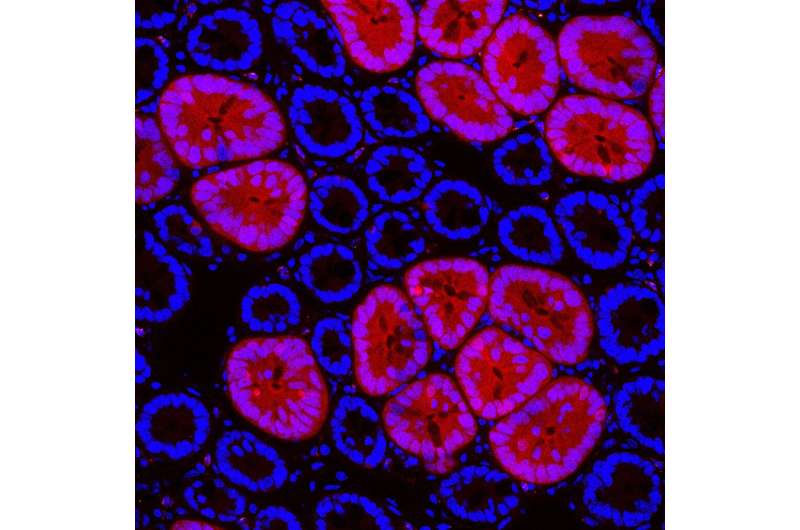
Oncogenic factors and ruthless crosstalk mechanisms
Using Red2Onco, the team was able to study the mechanisms initiated by two separate "first oncogenic hits." These are mutations to known proto-oncogenes, KRAS and PI3K, respectively, being known as pro-cancerous factors driving malignancy in declared tumors. To their surprise, the researchers found that even in the event of such a first oncogenic hit, the mutated pre-cancerous cell or "seeding cell" imparts a "negative influence" on its neighbors.
The surrounding normal tissue loses its stem cells, which in turn favors the territorial expansion of oncogenic mutant stem cells and their progenies. "Through this process of 'field transformation," the colonization of intestinal tissue by mutant cells increases the chance for further oncogenic hits, which may lead in turn to cancer," explains Benjamin Simons of the Gurdon Institute, University of Cambridge.
The first author on the study, Min Kyu Yum of the Gurdon Institute, expands: "Oncogenic mutant cells influence the fate behavior of their wild-type neighbors both directly, through secretion of signaling factors, and indirectly through induced changes in the shared tissue environment," before co-first author Seungmin Han sums up: "Using comparative single-cell analyses and organoid culture methods, we were able to dissect the molecular mechanisms that mediate cellular crosstalk."
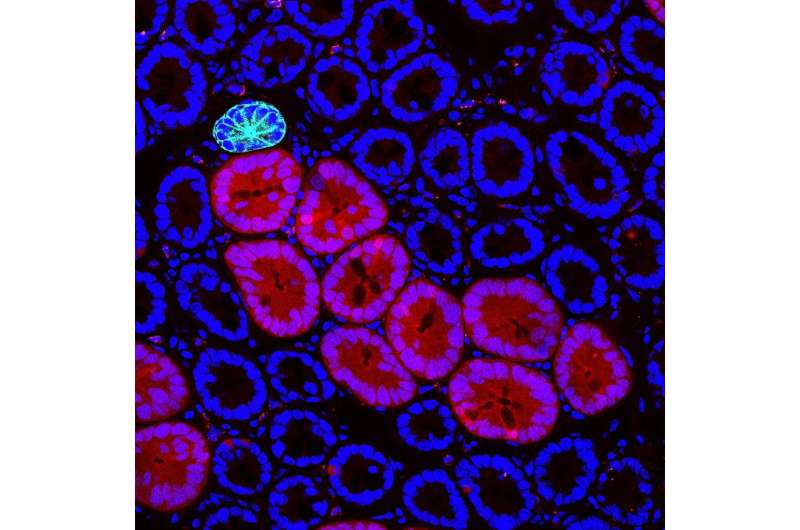
The aggressive influence can be suppressed
The relevance of the work is major, to say the least. In this sense, the authors have shown that inhibiting BMP (Bone Morphogenic Protein) signaling coming from the oncogenic mutant relieves the "negative influence" on the normal stem cells. "As well as enabling detection of early events of tumorigenesis, our discovery paves the way for intervention strategies that target cellular crosstalk mechanisms," concludes Min Kyu Yum.
More information: Tracing oncogene-driven remodelling of the intestinal stem cell niche, Nature (2021). DOI: 10.1038/s41586-021-03605-0 , www.nature.com/articles/s41586-021-03605-0