Breakthrough in understanding genesis of fibroids
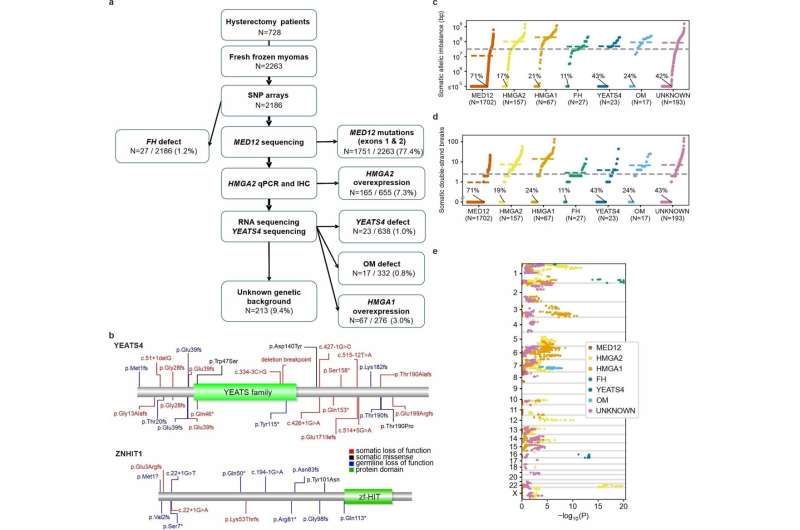
Scientists at the University of Helsinki and Helsinki University Hospital have made a breakthrough in understanding the genesis of uterine leiomyomas, also called fibroids.
Fibroids are extremely common tumors. They are a major burden for women's health worldwide, and the most common cause of hysterectomy. The Finland Myoma Study published in Nature found that the part of the human genome that controls expression of genes, is of major importance in fibroid development.
The findings of the new study represent a significant advance in fibroids research. Without detailed knowledge on the mechanisms of tumorigenesis involved, it would be difficult to develop targeted therapies for this condition affecting hundreds of millions of women.
Genes that are poised to change expression level are important for fibroid development
The researchers discovered that multiple tumors carried mutations in genes that were involved in trafficking certain type of histones, proteins that are important for the structure and functional properties of the genome. They next found that mutations in these same genes were important in hereditary predisposition to the disease.
The work also documented the many changes in the regulatory genome that these mutations exerted, and finally showed a strong effect on gene expression levels.
"In particular, genes that are poised to frequently turn on and off seem affected in fibroids," says Academy Professor Lauri Aaltonen.
This might explain why this new mechanism of tumorigenesis frequently affects the uterine muscle wall but rarely other tissue types, as the uterus needs to adjust to many changing external cues such as those governing the menstrual cycle and pregnancy.
"Thus, disturbances in genes that need to be poised to change expression level might harm the uterus more easily than other organs," explains Aaltonen.
"Some of the overexpressed genes might provide clues for development of new fibroid treatment options," Aaltonen says.
More information: Davide G. Berta et al, Deficient H2A.Z deposition is associated with genesis of uterine leiomyoma, Nature (2021). DOI: 10.1038/s41586-021-03747-1