Using AI for accurately predicting synergistic cancer drug combinations
A research team led by a scholar from City University of Hong Kong (CityU) has recently developed a novel artificial intelligence (AI) framework that can make synergistic anti-cancer drug combination predictions on both the therapeutic and toxic effects. The study revealed the potential of applying AI and machine learning to discover effective combinations in therapies for cancer and other complex diseases.
The research team was led by Dr. Zhang Qingpeng, Associate Professor from CityU's School of Data Science. The findings have been published in the Journal of the American Medical Informatics Association (JAMIA).
Applying AI in cancer therapies
To achieve the best therapeutic effect, multiple types of drugs are commonly prescribed by doctors to treat complex diseases like cancers and HIV/AIDS. However, the knowledge of the best combinations of drugs is usually based on clinical experience and judgements.
In recent years, AI and data science have become increasingly important in cancer treatment and drug discovery because of the capability to process large-scale complex biological and clinical data. As their first attempt at using AI in cancer drug discovery, Dr. Zhang's research group, whose research focus is data science, successfully developed "Graph Convolutional Network for Drug Synergy (GraphSynergy)". It is a novel AI framework that can make synergistic chemotherapy drug combination predictions.
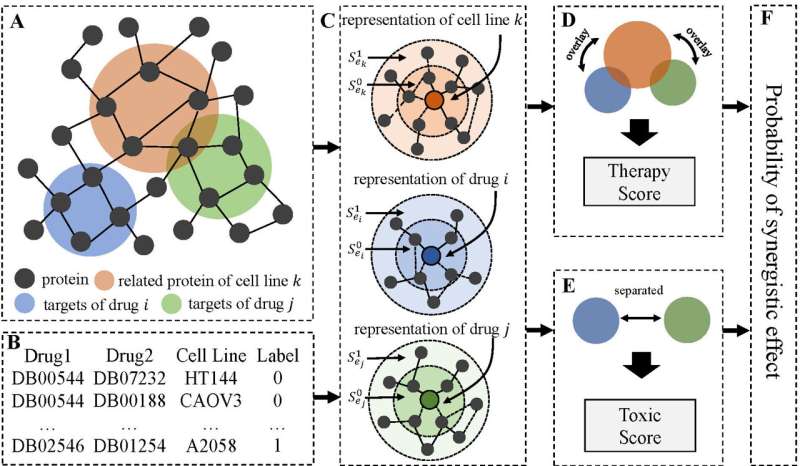
By analyzing the relations between the protein modules targeted by drugs and the protein modules associated with cancer cell lines in the human protein-protein-interaction network (PPI), and the interrelations among the protein modules, this deep-learning-based algorithm can identify synergistic drug combinations that yield effective therapy with less toxicity.
Analyzing the human protein-protein-interaction network
Dr. Zhang explained that the human body is a complex system, in which proteins in cells have numerous interactions and form a complex PPI network. Cancer cells are usually associated with multiple proteins, which have direct and indirect interrelations among each other and with other proteins in the PPI network. "How to mix and match multiple drugs to treat cancer is therefore very challenging. considering the complexity of the PPI network. So, we used AI to analyze the proteins that are targeted by drugs and the cancer cells, as well as the interrelations among the proteins, to help predict synergistic cancer drug combinations accurately," said Dr. Zhang.
He emphasized that the predictions are made by examining not only the directly targeted proteins by drugs and the cancer cells, but also the relations between the proteins, thus yielding a superior performance in identifying the best synergistic anti-cancer drug combination of low toxicity.
The research team compared the prediction performance of GraphSynergy with conventional prediction models and the latest drug combination prediction algorithms such as "DeepSynergy". They found that GraphSynergy's prediction performance is better in identifying synergistic drug combinations of low toxicity.
To verify the predictions made by GraphSynergy, the team selected clinically verified synergistic drug combinations to run the algorithm. And GraphSynergy predicted such combination is synergistic too.
According to the public dataset that the team obtained, the therapeutic effect of one of the drug combinations is considered not so significant. However, GraphSynergy predicted it as pharmacologically effective. The team further investigated that drug combination by reviewing the literature. It turned out that its effectiveness is recently verified in a clinical trial study. Dr. Zhang believed that this illustrated the forecast and prediction capability of AI algorithms.
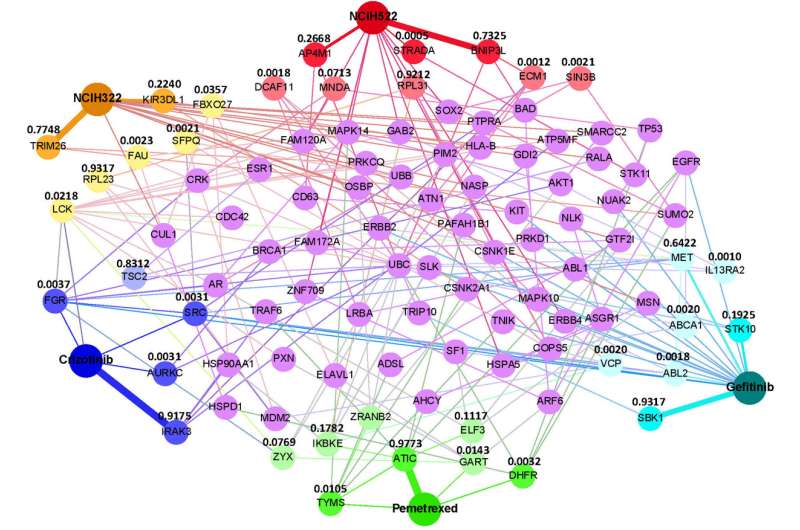
Dr. Zhang added that GraphSynergy could identify the pivotal proteins that play a role in making the cancer drugs effective. Such insight helps healthcare professionals to understand the predictions and provide guidance in future drug development.
Co-incubated by HK Tech 300 and HKSTP
"Our AI framework has the potential of optimizing cancer treatment at a low cost. We hope that it can provide grounds for doctors and patients to make medical decisions. Our next step will be applying the framework in drug development for cancer and other diseases," concluded Dr. Zhang.
To explore the translation of their research into an actual application, core members of the research team has established "COMPASS Health Technology", which is one of the start-up teams co-incubated by HK Tech 300, CityU's flagship innovation and entrepreneurship program, and Hong Kong Science and Technology Parks Corporation (HKSTP).
More information: Jiannan Yang et al, GraphSynergy: a network-inspired deep learning model for anticancer drug combination prediction, Journal of the American Medical Informatics Association (2021). DOI: 10.1093/jamia/ocab162