Type 1 diabetes can be predicted with epigenetic changes
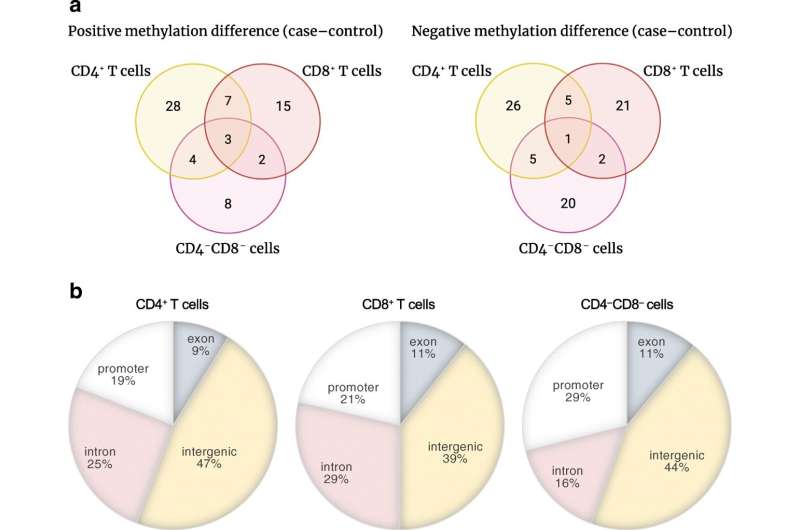
Children who develop type 1 diabetes show epigenetic changes in the cells of their immune system already before the antibodies of the disease are detected in their blood. The findings of two new studies offer new opportunities to identify the children with the genetic risk for developing diabetes very early on.
Epigenetic changes can affect how our genes work. Environmental factors, such as viral infections, can cause epigenetic changes.
The findings in the epigenetic makeup linked to diabetes were discovered in two new studies led by researchers from Turku Bioscience at the University of Turku, Finland.
"We uncovered previously unknown, early-onset epigenetic changes. They offer us new opportunities to further develop ways to identify children who have a risk of developing type 1 diabetes even before they get sick," says Professor Riitta Lahesmaa, Director of Turku Bioscience and a group leader in the InFLAMES research flagship initiative.
Earlier studies have shown that certain antibodies detected in children's blood samples indicate an increased risk of developing type 1 diabetes in the near future. So that medical professionals could intervene in the disease even sooner, earlier disease indicators than the antibodies are needed to detect the risk. This involves searching for biomarkers indicating type 1 diabetes, and epigenetic changes could be such a biomarker.
"Our observations on epigenetics are extremely important as our goal is to develop methods and tools to prevent the onset of type 1 diabetes in children who are at risk of developing the disease," says Professor Laura Elo. Elo is the Director of the Medical Bioinformatics Centre at Turku Bioscience and a group leader in the InFLAMES research flagship.
Finnish children have an increased risk of developing type 1 diabetes
In Finland, children's risk of developing type 1 diabetes is the highest in the world. In addition to the genetic susceptibility, environmental factors have a great significance for developing the disease. The environmental factors include, for example, excessive level of hygiene, biodiversity loss, and environmental toxins.
The newly published studies are based on long-term interdisciplinary research collaboration with international partners. The project has included doctors who are in charge of the patients and also conduct clinical research, researchers in molecular medicine and immunology, and experts in computational science. In the studies, researchers analyzed longitudinal samples with deep sequencing covering the entire genome as well as with computational methods and artificial intelligence.
More information: Inna Starskaia et al, Early DNA methylation changes in children developing beta cell autoimmunity at a young age, Diabetologia (2022). DOI: 10.1007/s00125-022-05657-x
Tomi Suomi et al, Type 1 Diabetes in Children With Genetic Risk May Be Predicted Very Early With a Blood miRNA, Diabetes Care (2022). DOI: 10.2337/dc21-2120