Multi-omics modeling in a twin cohort to predict blood pressure values

Based on data from more than 400 twins, researchers used multi-omics data integration to develop a model for predicting blood pressure values. Their research is described in the peer-reviewed OMICS: A Journal of Integrative Biology.
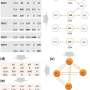
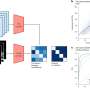
The investigators utilized a multi-omics regression-based method called sparse multi-block partial least square to predict systolic and diastolic blood pressure values. The study of high blood pressure has high public health importance, as it is known to increase the risk of cardiovascular, cerebrovascular and renal disease, amongst other common chronic human diseases. The authors, Gabin Drouard, Miina Ollikainen, and Jaakko Kaprio from the Institute for Molecular Medicine Finland, University of Helsinki, and coauthors in Finland and Augusta, Georgia, integrated blocks of omics—including transcriptomic, methylation, and metabolomic data—as well as polygenic risk scores into the modeling.
"In addition to revealing interesting inter-omics associations, we found that each block of omics heterogeneously improved the predictions of blood pressure values once the multi-omics data were integrated," stated the investigators.
"Hypertension is a high-prevalence multi-factorial disease. The new study by Drouard and colleagues is important for two reasons. First, it presents a multi-omics approach to integrate single omics analyses. Second, it illustrates the exploratory and predictive gains achieved by multi-omics study of a complex phenotype such as blood pressure. I believe this study is significant in the current era of systems medicine and planetary health," says Vural Özdemir, MD, Ph.D., DABCP, Editor-in-Chief of OMICS.
More information: Gabin Drouard et al, Multi-Omics Integration in a Twin Cohort and Predictive Modeling of Blood Pressure Values, OMICS: A Journal of Integrative Biology (2022). DOI: 10.1089/omi.2021.0201