July 29, 2022 report
Enzyme linked to most prevalent mutational signatures in cancer cell lines
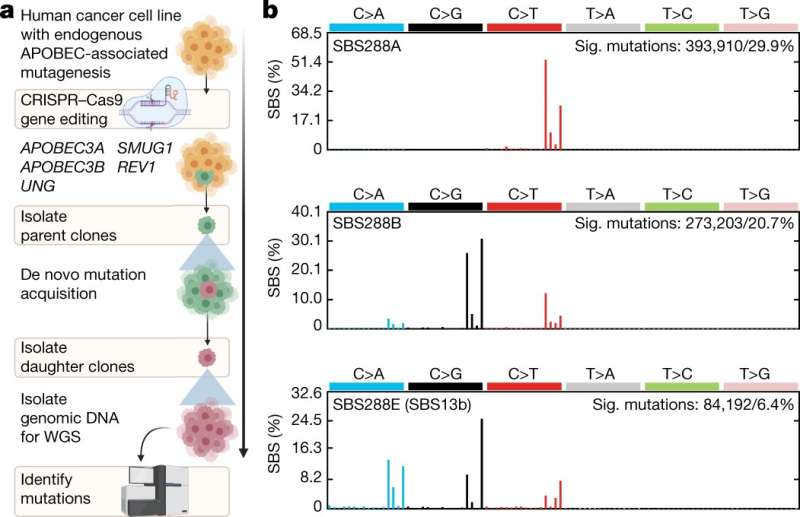
A team of researchers affiliated with several institutions in the U.S. and the U.K. has found a link between the APOBEC3A enzyme and the most prevalent mutational signatures in cancer cell lines. In their paper published in the journal Nature, the group describes removing APOBEC3 mutations from human cancer cell lines to learn more about possible clustered signatures.
Prior research has shown that the majority of cancer genomes have mutational signatures that can be associated with a family of enzymes known as APOBEC3. This work has suggested that these enzymes play a role in mutagenesis. Other prior work has suggested that the specific enzyme APOBEC3B is the most likely member of the family behind the ability of cancer cells to mutate. In this new effort, the researchers have taken a closer look at another member of the family—APOBEC3A—and have found evidence that it is actually the enzyme most responsible for activation of mutations.
To learn more about possible APOBEC3 signatures, the researchers removed the mutations that gave rise to differences in the enzyme family, namely APOBEC3A and APOBEC3B, from cells making up cancerous tissue in multiple human cell line samples. In all, they worked with 251 tissue samples taken from patients with bladder, breast or lymphoma cancer.
The researchers then analyzed the samples to identify the signatures that were missing. They discovered that APOBEC3A had a stronger signature presence than APOBEC3B, suggesting that it is more likely the enzyme that is responsible for instigating cancerous mutations. They also note that their work is the first to show evidence of endogenous APOBEC enzymes as a main driver of mutational signatures in cancer cells.
The researchers note that their work did not rule out APOBEC3B as a contributor to mutational activity, however, because in some cells lines, deletion of APOBEC3B actually resulted in an increase in mutations. This finding suggested that the two enzymes may play different roles in different cells. They acknowledge that more work is required to discover whether such roles are mutually exclusive or if the enzymes work together. The researchers conclude that their work also backs up theories two decades in the making that surround the role played by the APOBEC3 enzyme family.
More information: Mia Petljak et al, Mechanisms of APOBEC3 mutagenesis in human cancer cells, Nature (2022). DOI: 10.1038/s41586-022-04972-y
© 2022 Science X Network
















