Innovative screening method for medical agents based on genome sequence data
A team of scientists led by Professor Dr. Yvonne Mast, head of the Department of Bioresources for Bioeconomy and Health Research at the Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures in Braunschweig, Lower Saxony, has developed a new screening strategy for active medical substances. This strategy is based exclusively on analyzing genome sequence data.
Summarizing the advantages of her approach, Mast explains, "The advantage of our prediction strategy is that it can be used to screen purely digitally as well as very efficiently for specific potential drug producers and, in some cases, to screen out bacterial strains that are most likely to produce known substances. This makes the process of analyzing and finding active ingredients much easier. This new methodology is another important contribution in the rapidly developing research field of genome mining for the discovery of new active ingredients."
In recent years, fewer and fewer new active ingredients, such as antibiotics, have been discovered and developed for medical use, while simultaneously the threat of multidrug-resistant pathogens have been rapidly increasing worldwide. A major problem in medical drug research is the high recovery rate of already known substances, which makes conventional screening methods highly inefficient.
Digitization in the field of active ingredient research
Digital sequence information is gaining more and more importance in active ingredient research. The team led by microbiologist Prof. Dr. Yvonne Mast has developed a prediction strategy based on genome sequence information from bacterial active ingredient producers that makes it possible to predict the production of a specific group of active ingredients, the so-called protein synthesis inhibitors. These active ingredients target the bacterial ribosome, which is an important place of action for many important antibiotics like erythromycin, tetracycline and clindamycin.
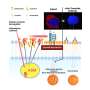
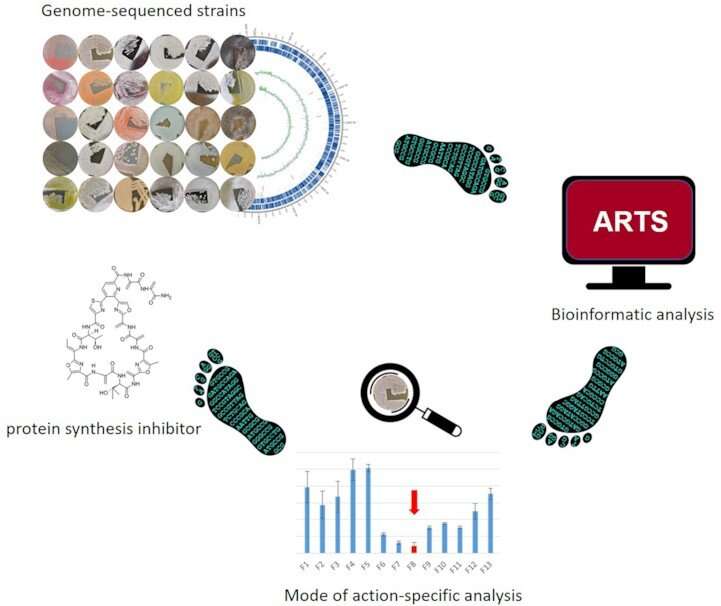
The Braunschweig-based team of scientists was able to derive their prediction strategy by identifying specific indicator genes in the genome of drug producers from the bacterial group of Actinomycetes, which indicates protein biosynthesis inhibitor synthesis performance. The scientists provided proof of function by identifying the protein biosynthesis inhibitor amicoumacin from various Actinomyces strains of the DSMZ bacterial collection. The methodology has been named Ψ-footprinting and was recently published in NAR Genomics and Bioinformatics.
More information: Franziska Handel et al, Ψ-Footprinting approach for the identification of protein synthesis inhibitor producers, NAR Genomics and Bioinformatics (2022). DOI: 10.1093/nargab/lqac055