Study investigates how antiviral treatment promotes emergence of new SARS-CoV-2 variants
Do patients with prolonged infections contribute to the emergence of new SARS-CoV-2 variants? A research team from the Leibniz Institute of Virology (LIV) and the University Medical Center Hamburg-Eppendorf (UKE) has investigated this question in more detail and has now been able to show that it is not the prolonged infection per se that leads to the emergence of new variants. Rather, it requires an evolutionary bottleneck such as can be created by antiviral treatment. The results have now appeared in the journal Cell Reports Medicine as a pre-proof version.
Prolonged SARS-CoV-2 infections occur primarily in immunocompromised patients and have been repeatedly discussed as important contributors in viral evolution: Reduced immune restriction could lead to a broad increase in viral diversity within the host, favoring the emergence of new variants, especially if antiviral treatments—such as with Remdesivir or convalescent plasma—exert selection pressure for the acquisition of escape mutations.
Analysis of genomic diversity in prolonged infections
In the published study, a research team led by Prof. Adam Grundhoff (LIV) and Prof. Nicole Fischer (UKE) has now investigated whether patients with prolonged infections generally exhibit increased viral evolution, which could allow for the more rapid emergence of SARS-CoV-2 variants, or whether certain treatment regimens promote the emergence of new mutations.
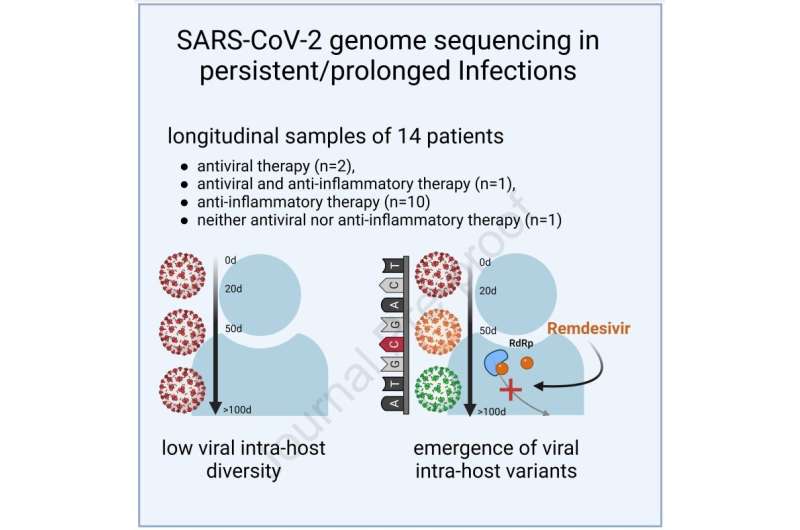
For this purpose, the genomic diversity within the host was investigated by whole genome sequencing in longitudinal samples from 14 patients with prolonged viral persistence (30–146 days) during severe COVID-19 disease; including immunocompromised and immunocompetent patients with or without antiviral treatment to assess the occurrence of mutations with and without selection pressure.
"Overall, the virus was remarkably stable in the vast majority of individuals studied. However, in one patient treated with Remdesivir, we observed that a high number of mutations occurred immediately after the start of treatment—including at least one mutation that was highly likely to confer increased resistance to Remdesivir," explains Prof. Adam Grundhoff, head of the LIV Virus Genomics Research Group.
Antiviral treatment promotes evolutionary bottleneck
Patients with prolonged SARS-CoV-2 infection and antiviral Remdesivir treatment showed a significant increase in viral intra-host diversity with newly emerging mutations. In contrast, in patients receiving anti-inflammatory treatment alone, the emergence of new variants was observed only sporadically.
"Our work shows that it is not the long duration of the infection per se that leads to the emergence of new variants. Rather, this requires an evolutionary bottleneck, such as can be created by antiviral treatment. This finding is particularly important in view of the recent discussions on the use of Remdesivir for the treatment of non-hospitalized high-risk patients, but also for the introduction of potentially new antiviral therapeutics," says Prof. Nicole Fischer from UKE.
More information: Andreas Heyer et al, Remdesivir-induced emergence of SARS-CoV2 variants in patients with prolonged infection, Cell Reports Medicine (2022). DOI: 10.1016/j.xcrm.2022.100735