Single clinical test provides more answers about COVID
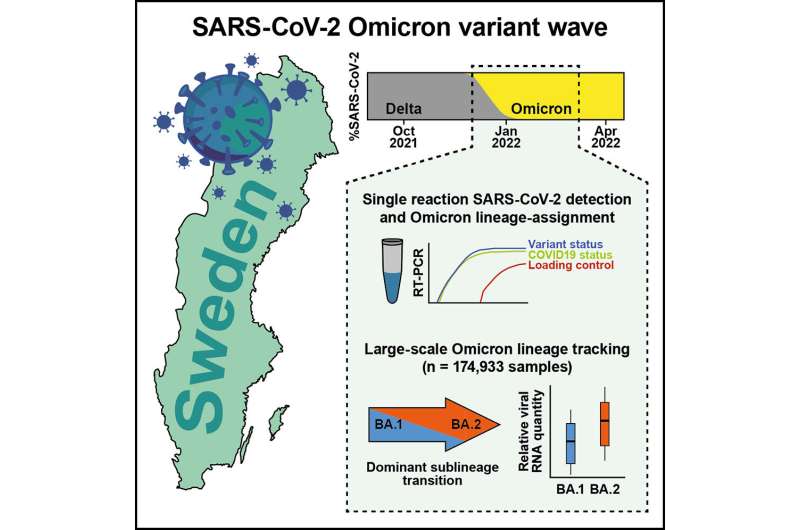
Researchers at Karolinska Institutet present the results of a refined clinical COVID-19 test, which has been used to track the spread of the omicron variant in real time in the Swedish population. The study, published in the journal Med, provides new insights into the dominance transition of omicron sublineages that occurred consistently across the world.
During the first wave of the COVID-19 pandemic in the spring of 2020, Björn Reinius' research group at Karolinska Institutet developed a fast, inexpensive and accurate COVID-19 clinical analysis, which helped to enable large-scale clinical testing even in the early stages of the pandemic.
The method was subsequently used in millions of clinical COVID-19 tests in Sweden and across the globe. The research group is now publishing the results of a refined analysis method.
"In response to the rapid spread of a new COVID variant, omicron BA.1, at the beginning of December last year, we quickly worked to refine the method further so that we could also classify which variant of the COVID-19 virus an infected person was carrying," says Antonio Lentini, researcher at the Department of Medical Biochemistry and Biophysics, Karolinska Institutet, and the study's first author.
The new method, a custom variant typing RT-PCR assay, made it possible to tell whether a patient sample was positive for COVID-19 and whether the infection was the of omicron BA.1 variant.
"Previously, we have had to perform sequencing or several separate PCR assays in addition to the primary clinical test to obtain these results. During the omicron wave at the turn of the year 2021/2022, however, the laboratories were completely overburdened by primary testing, and this method helped us track the development of the omicron variant in the Swedish population in real time," says Björn Reinius, research group leader at the same department and the study's last author.
According to the researchers, the new method is both accurate and reliable. Quality checks were performed against viral genome sequencing, showing that the method had greater than 99% concordance in variant classification.
Tracking the rapid shift from omicron BA.1 to BA.2
Just a few weeks into the omicron wave, an even more infectious form of the virus arrived in Sweden—omicron BA.2. The method developed by the researchers proved to be useful in this case as well. If SARS-CoV-2 was detected in the assay, but not BA.1, the test was positive for BA.2 because other COVID variants had already been outcompeted.
Between January and March of 2022, the research team analyzed 174,933 clinical upper respiratory samples in close collaboration with ABC Labs, which was assisting the Public Health Agency of Sweden and Swedish health care regions with analysis capacity at the time.
"We saw that virus levels, quantified as viral genome copies, in the upper respiratory tract of people infected with omicron BA.2 were almost twice as high compared to BA.1 cases. Our data can help explain why omicron BA.2 spread so aggressively and quickly pushed BA.1 out of the population," says Björn Reinius.
The researchers are now working to develop new methods for diagnosing viral and bacterial infections. Their goal is to develop a single clinical assay that can detect a large number of infectious diseases simultaneously.
More information: Antonio Lentini et al, Monitoring of the SARS-CoV-2 Omicron BA.1/BA.2 lineage transition in the Swedish population reveals increased viral RNA levels in BA.2 cases, Med (2022). DOI: 10.1016/j.medj.2022.07.007