New machine learning tool can help predict patients most at risk of COVID-19
Researchers have created a machine learning tool that can help identify patients who are most at risk of developing COVID-19 while in hospital.
The tool, which is a form of artificial intelligence (AI), was able to predict patients at high risk of developing COVID-19 with 87 percent accuracy in a study. The technology has been developed by researchers at Imperial College London and the Infection Prevention and Control (IPC) unit at Imperial College Healthcare NHS Trust.
Researchers developed the tool using routine hospital and patient data. They trained it to identify risk factors associated with COVID-19 infections such as age, gender, contacts with other infectious patients, where beds were situated and how patients moved around the hospital.
Predicting which patients are at risk of developing infection while in hospital can help prevent onward transmission to other patients and staff.
The tool was tested using data from inpatients admitted to Imperial College Healthcare hospitals during the first two waves of COVID-19 infection and validated using data from Geneva University Hospitals. The study, published in The Lancet Digital Health, is the first to use patient contact information to accurately predict the risk of developing COVID-19 in hospital.
The study used data from Imperial College Healthcare's Clinical Analytics, Research and Evaluation (iCARE) environment, which provides access to routinely collected, anonymized healthcare data for research for direct patient benefit.
Surveillance tool
The team believes this tool could be applied to other infections that some patients can be at risk of developing whilst in hospital, such as Clostridium difficile (C difficile)—a type of bacteria that can cause diarrhea.
Lead author of the study Ashleigh Myall from the Department of Mathematics at Imperial College London, says that "throughout the COVID-19 pandemic some patients developed the infection during their hospital stay. There is a need to develop predictive models to identify patients who are most at risk of contracting COVID-19 and intervene to mitigate against poor patient outcomes. We can do this alongside the usual measures to minimize outbreaks and further transmissions."
"We have designed a machine learning framework that can identify patients who are most at risk of catching COVID-19 with up to 87 percent accuracy. This framework could be used as part of a range of surveillance tools to enhance infection, prevention and control strategies, especially during the winter months when COVID-19 infections spread more easily."
"The study does have its limitations as the training and testing period occurred before the UK's vaccination rollout, so it does not take into account an individual's vaccination status in determining risk. However, our model is highly predictive and could be used for other infectious viruses."
Sid Mookerjee, co-author of the study and operational lead for antimicrobial stewardship, surveillance and epidemiology at Imperial College Healthcare NHS Trust, says that "the COVID-19 pandemic has energized researchers and clinicians to invest time, funds and effort to respond to current and future pandemics."
"Accurately predicting patients' risk of developing infections such as COVID-19, C difficile and other infectious diseases, is a much sought-after clinical solution."
"Through this work we go some way in showcasing a new and highly predictive approach to identifying patients at risk of developing infections whilst seeking hospital care. This can help hospital managers and clinicians design safe and effective patient care pathways and bed management, helping deliver world class care."
Details of the study
Transmission of COVID-19 infections within healthcare settings has been well documented. COVID-19 infections that have developed after admission to hospital have been reported to account for 12–15 percent of all COVID-19 cases in healthcare settings and up to 16.2 percent at the peaks of the pandemic.
Traditionally, prediction of infections that can develop in the healthcare setting relies on identifying risk factors such as age, gender identity, comorbidities and patients' length of stay and has not taken into account patient contacts, locations or patient-flow through the hospitals.
Although these approaches alone can perform reasonably well in identifying predictive risk factors of HCAIs they overlook the fact that HCAIs' spread depends largely on the patient's contacts, which can vary.
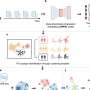
The team wanted to see whether a machine learning tool could predict patient risk of HOCI using patient contact data. The team combined patient contacts data based on bed allocation with clinical and hospital data from the iCARE system, which is supported by the NIHR Imperial BRC, into a forecasting framework to predict patient risk of HOCIs.
HOCIs are defined as infections in patients with a positive SARS-CoV-2 test—a type of the coronavirus that causes COVID-19—three or more days after admission. Patient contacts were defined as patients coinciding on the same day in the same room, ward and building, regardless of COVID-19 prevention strategies such as environmental ventilation and PPE use.
Testing
The tool could not account for the mitigating effect of standard national IPC measures such as PPE usage, ventilation, hand hygiene and cleaning in reducing the risk of patients developing COVID-19. The study was also conducted before the wider national roll out of COVID-19 vaccination, a significant factor in reducing risk of developing COVID-19.
The framework works by analyzing risk factors associated with COVID-19 infections and hospital data of patients and then gives a predicted risk score between 0 and 1.
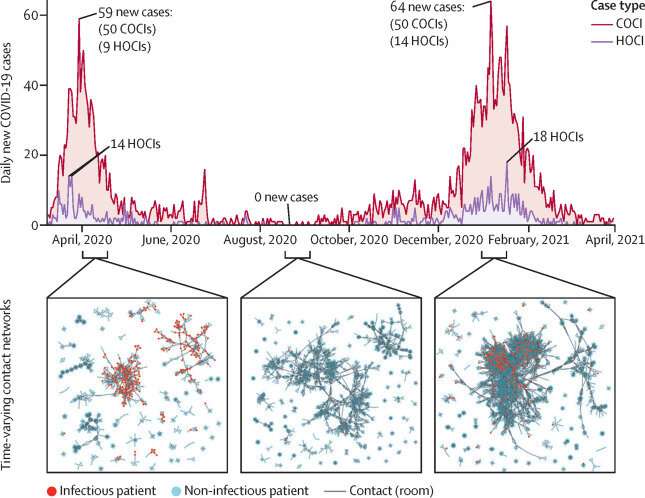
The model was tested using data from 51,157 patients at Imperial College Healthcare NHS Trust hospitals during the first two UK surges of COVID-19 (March to May 2020 and September 2020 to April 2021). A total of 3749 patients tested positive for COVID-19 three or more days after their admission to hospital during this time, 87 percent of which were accurately predicted by the machine learning framework.
The team compared this with a control group who did not test positive for COVID-19. They then validated the framework by applying it to data from 2021 external dataset from Geneva University Hospitals.
The researchers will carry out further work to extend the framework to include the omicron strain of COVID-19, other infectious diseases and understand how the framework could be integrated into existing IPC guidelines.
More information: Ashleigh Myall et al, Prediction of hospital-onset COVID-19 infections using dynamic networks of patient contact: an international retrospective cohort study, The Lancet Digital Health (2022). DOI: 10.1016/S2589-7500(22)00093-0