February 22, 2023 feature
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Identifying drug target candidates to treat pediatric rhabdomyosarcoma tumors
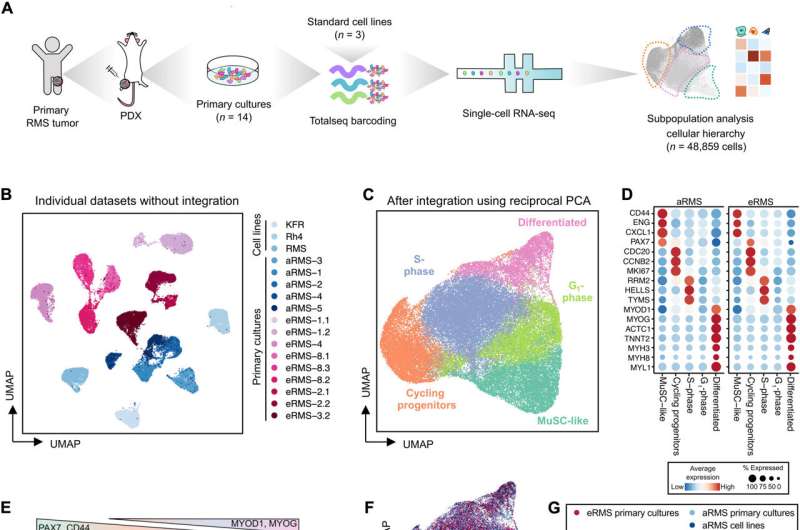
Cancer biologists are yet to understand the mechanisms and cellular hierarchy leading to the developmental arrest in rhabdomyosarcoma (RMS)—a group of pediatric cancers, which remain enigmatic.
In a new report in Science Advances, Sara G. Danielli and an international, interdisciplinary team of scientists at the department of Oncology and Children's Research, Stem Cell Biology, and Translational Research in Child and Adolescent Cancer, in the U.S., Switzerland, France, Germany and Spain, combined a myriad of biological methods to understand the etiology and cellular basis of RMS oncogenesis.
The team incorporated single-cell RNA sequencing, mass cytometry and high-content imaging methods to understand the intra-tumoral diversity of patient-derived biopsies. The aggressive alveolar subtype (aRMS) contained plastic muscle stem-like cells and precursors of tumor growth alongside a subpopulation of differentiated cells without proliferative potential that led to better outcomes.
The scientists implemented chemotherapy and observed the dynamics of the aggressive alveolar version of the disease. They then screened the drug candidates and their capacity to propagate the disease towards clinically favorable sub-populations and identified a combination of rapidly accelerated fibrosarcoma (RAF) and mitogen activated protein kinase (MEK) inhibitors to induce muscle differentiation and inhibit tumor growth. These outcomes provide insights to develop biological states underlying the disease, including aggressiveness, chemoresistance and cell growth, while identifying the Ras pathway as a promising therapeutic target.
Rhabdomyosarcoma (RMS)
Childhood cancer leads to dysregulated human development and is a leading cause of disease-related morbidity and mortality in children and adolescents. Recent advances in single-cell technologies can assist the characterization of intratumoral diversity and shed light to the phenotypic plasticity across several cancer types to highlight their role as emerging hallmarks of oncogenesis.
Researchers are studying the developmental hierarchies of childhood disease to identify their origin and develop effective treatment strategies that target the cellular components. Rhabdomyosarcoma is a common pediatric soft tissue sarcoma occurring as embryonic and alveolar (aRMS) subtypes, of which the latter is more aggressive due to its underlying genetic basis.
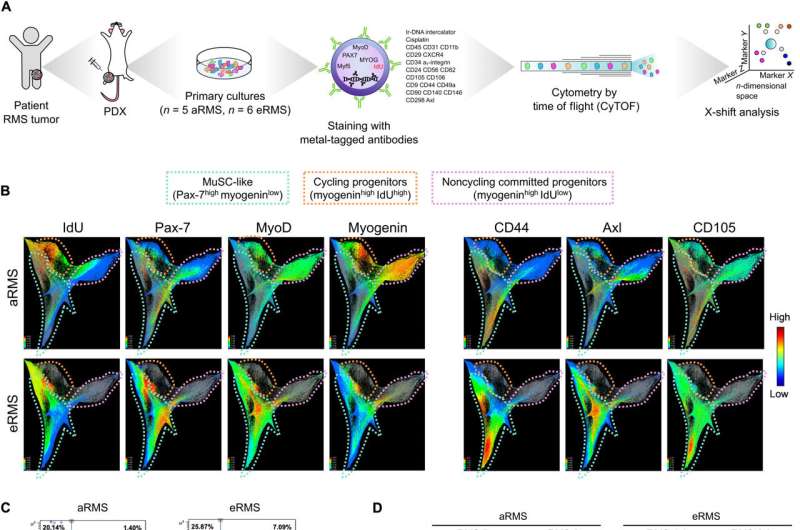
In the current study, Danielli and colleagues combined scRNA sequencing, mass cytometry, and high-content imaging analysis to examine the intratumoral heterogeneity of the cancer cell lines and of primary cultures obtained from patient-derived xenografts.
The team then screened regulators of the alveolar rhabdomyosarcoma (aRMS) cell fate via a library of pharmaceutical compounds, including RAS pathway inhibitors trametinib with dabrafenib or regorafenib to direct these cancer cells to differentiate. The combinatorial therapies potentially suppressed the growth of the tumor in patient-derived xenografts to provide a strong rationale to clinically manipulate the RAS pathway to counter oncogenesis in patients.
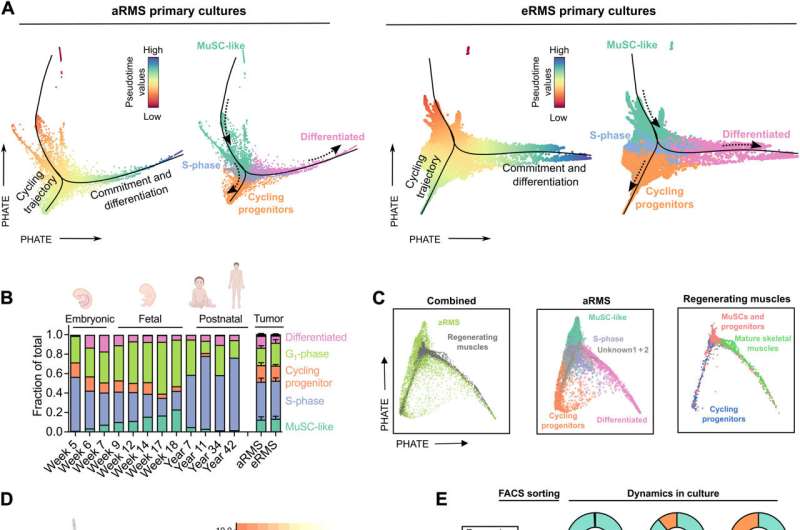
Using single-cell RNA sequencing to identify the distinct muscle development states
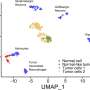
The researchers used droplet-based single cell RNA sequencing to understand the intratumoral diversity of cancer cells by profiling 14 patient-derived xenografts and three conventional alveolar rhabdomyosarcoma cell lines in comparison with preceding studies. The team increased the interpatient variability by choosing disease models that originated from diverse oncogenic subtypes, including primary and metastatic sites among diagnostic and recurrent patients who had or had not undergone pretreatment.
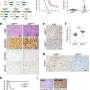
The outcomes revealed how the RMS tumors contained myogenic cells stalled in an immature transcriptional state to ultimately form a minority of differentiated cells. The scientists confirmed the presence of the subpopulations at the protein level by staining the primary cell cultures with an isotope-conjugated antibody panel to isolate muscle stem-like cells, which they identified using single-cell RNA sequencing to define the distinct states of myogenesis.
Next, they profiled the cells by using cytometry via time-of-flight analyses, where the research outcomes indicated the aRMS cell subset to be of specific interest. The single-cell protein analysis outcomes aligned with the transcriptomic analysis.
Mechanism-of-action of the cancer phenotype and therapeutic intervention in the lab
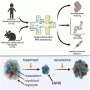
The team demonstrated the mechanism-of-action of the RMS cancer phenotypes that mirrored the cell-fate decisions of developing, healthy and regenerating skeletal muscle stem-like cells. They studied this relationship via a trajectory inference model based on Slingshot and PHATE data visualization tools, to envision the structure and transitions in high-dimensional biological data. When the scientists projected the single-cell RNA sequencing atlas of the developing human skeletal muscles to the oncogenic single-cell transcriptome; the tumor cells mostly mapped onto muscle cells that were transiting from the embryonic to the fetal stages.
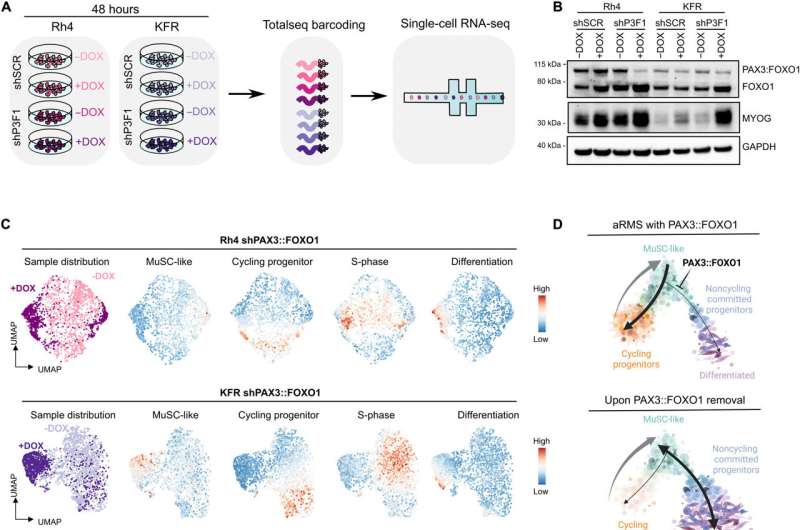
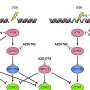
The scientists noted the major driver of the aRMS subtype to be the PAX3::FOX01 fusion gene; known to repress myogenic differentiation. To study the genes, the team used two knockdown aRMS cell lines against the fusion genes; these treatment measures resulted in decreased levels of the fusion gene to highlight the gene knockdown capacity of the cell lines to either undergo differentiation or halt in a muscle stem-like state, allowing continued targeted fusion-gene therapy. It also appeared that some cell lines that intrinsically resisted treatment underwent cellular trajectory rewiring.
Differentiation therapy with drug combinations
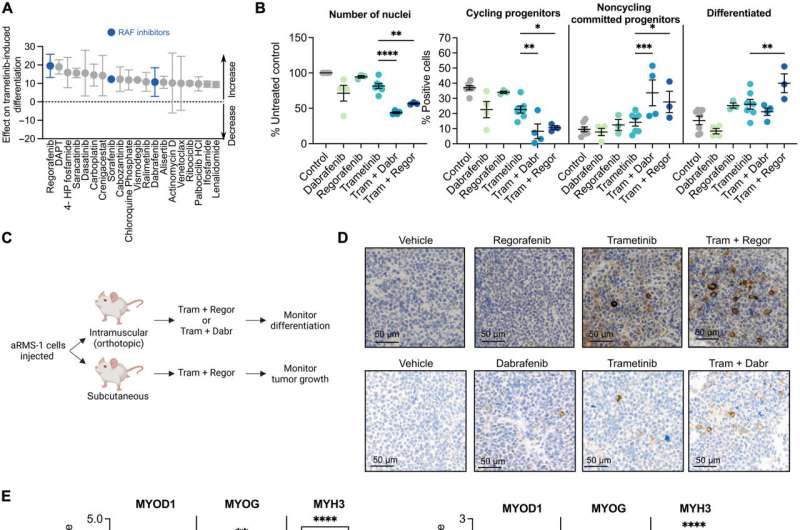
The researchers observed the mechanism-of-action of Trametinib, which hijacked the cell fate of the treatment resistant aRMS cell line and redirected these cells toward differentiation. Of the screened drug moieties Trametinib, Cobimetinib and Erdafitinib induced robust myogenic differentiation.
Since monotherapy remains to succeed in clinical trials, the team identified the more effective combinatorial drug screening to demonstrate the trametinib-induced suppression of the RAS pathway in combination with RAF inhibition as a feasible treatment strategy for aRMS.
Outlook
In this way, Sara G. Danielli and colleagues developed a comprehensive single-cell transcriptomic and proteomic atlas of rhabdomyosarcoma (RMS). The atlas detailed cellular and functional diversity of the disease and revealed key cellular and molecular signatures suited for therapeutic intervention to overcome chemoresistance and tumor relapse. The researchers described the mechanism-of-action of the cell fate underlying impaired differentiation in the aggressive alveolar rhabdomyosarcoma (aRMS) cancer subtype. The work sheds light on how to therapeutically restore myogenic differentiation and block tumor growth.
More information: Sara G. Danielli et al, Single-cell profiling of alveolar rhabdomyosarcoma reveals RAS pathway inhibitors as cell-fate hijackers with therapeutic relevance, Science Advances (2023). DOI: 10.1126/sciadv.ade9238
Simone Hettmer et al, Muscling in: Uncovering the origins of rhabdomyosarcoma, Nature Medicine (2010). DOI: 10.1038/nm0210-171
© 2023 Science X Network