May 19, 2023 report
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
A global effort to identify critical illness in some COVID-19 patients highlights genetic risk, potential treatments
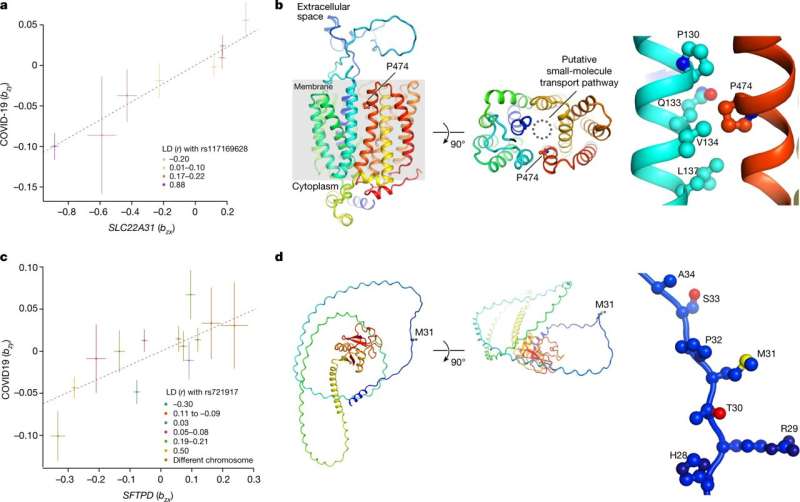
Researchers at the University of Edinburgh in the U.K. have led a study in collaboration with scientists worldwide, looking into cases of critical illness in COVID-19 patients.
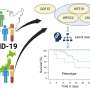
Critical illness in COVID-19 is an extreme and clinically consistent disease phenotype the team has found presenting in patients with shared genetic attributes. These shared genetics hint at a shared mechanism for the critical illness not seen in other patients and potential therapies to address the condition.
Patients with confirmed COVID-19 and requiring continuous cardiorespiratory monitoring or organ support (a generalizable definition for critical illness) were recruited in 2020–2022.
Researchers analyzed 24,202 cases of COVID-19 with critical illness with a combination of microarray genotyping and whole-genome sequencing data from the international GenOMICC study (11,440 cases) and other studies recruiting hospitalized patients with severe and critical illness, including the COVID-19 Human Genetics Initiative, the International Severe Acute Respiratory and Emerging Infection Consortium, the Spanish Coalition to Unlock Research on Host Genetics consortium and 23andMe.
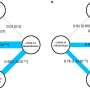
The team found 49 genome-wide significant associations, of which 16 have not been reported previously and 196 significantly associated genes in a gene-level analysis. Although the implicated variants are not directly causing illness in the patients, they can highlight molecular mechanisms that make some COVID-19 infections much more severe. The findings are published in the journal Nature.
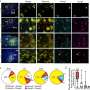
Many genes implicated in critical COVID-19 are highly expressed in the monocyte-macrophage system, which has poor coverage in existing expression quantitative trait loci datasets. Macrophages synthesize many substances involved in host defense and inflammation and play a pivotal role in immune system reactions.
Additionally, the investigation found variation in circulating protein levels with 15 unique proteins linked to critical illness and some with well-studied biomarkers that make them good candidates for drug targeting.
The research has identified several potential druggable targets in multiple systems, including inflammatory signaling, monocyte-macrophage activation and endothelial permeability. Some of the targets found have already seen positive results with therapeutic signals in multiple drug trials, providing a good proof-of-concept for drug target identification using comparative genetics.
More information: Erola Pairo-Castineira et al, GWAS and meta-analysis identifies 49 genetic variants underlying critical COVID-19, Nature (2023). DOI: 10.1038/s41586-023-06034-3
© 2023 Science X Network