This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
COVID-19 lessons learned: 'Why aren't we working on all diseases like this?'
Within months of COVID-19's discovery, UCSF Quantitative Biosciences Institute (QBI) and a group of international scientists charted the first roadmap to possible future treatments based on existing medications. Now, QBI reveals secrets to its success—and what they tell us about preparing for future pandemics.
UCSF was a leader in the response to COVID-19, caring for some of California's first COVID-19 patients, setting up innovative testing and vaccination programs and offering care to vulnerable populations in California and beyond.
QBI combines fields like chemistry, biology and physics to understand what fuels disease at a genetic or cellular level. The institute uses this information to find new ways to diagnose and treat illnesses. In some cases, this means breathing new life into old drugs.
As other researchers focused on developing COVID-19 vaccines, QBI turned its attention to possible treatments using existing drugs to stop or slow the virus. To do this, QBI director Nevan Krogan, Ph.D., formed the QBI Coronavirus Research Group to study how the virus attacks cells. QBI Chief Operating Officer Jacqueline Fabius coordinated the group's work as it swelled to include more than 120 scientists around the world.
Krogan and Fabius explain what COVID-19 taught them about doing science during a pandemic in a new editorial in the journal Cell Host & Microbe. We wanted to learn more.
Science is typically competitive. Why did COVID-19 require collaboration?
Krogan: It was clear we wouldn't be able to do everything alone. In early March 2020, for example, we had an initial list of proteins we thought played a role in COVID-19 infection. We also had a list of medicines, or drug compounds, that we thought might do something.
Fabius: But we didn't have the live virus to test our theories. Two of our partners did.
As travel bans loomed, we raced to ship compounds to the Icahn School of Medicine and Institut Pasteur—some of the only labs globally working with live coronavirus samples at the time. We even explored diplomatic routes for the compounds to reach Paris with San Francisco's French Consulate, given international travel freezes.
Within days, both labs had the shipments and began testing.
What impacts did you see from those collaborations?
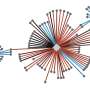
Krogan: Ultimately, QBI, UCSF-affiliated Gladstone Institutes (where Krogan is a senior investigator), France's Institut Pasteur and the Icahn School of Medicine at Mount Sinai in New York mapped more than 300 "doorways" that play a role in COVID-19 infection. They found about 70 existing or developmental medicines that could possibly protect cells from the viral invader. Some of these medicines included common allergy medications, antipsychotic drugs and antianxiety medicines.
One, a cancer drug, has shown early promise in reducing COVID-19 viral loads in clinical trials much like existing COVID-19 medications like Paxlovid. The less virus a person has in their body, the less sick they are likely to become. Scientists are hopeful it may also prove effective against future viruses similar to COVID-19.
Maps like the one we produced can take anywhere from two to three years. We produced our COVID-19 protein map in two weeks. The QBI Coronavirus Research Group eventually included hundreds of scientists working at breakneck speed—literally around the clock in shifts—seven days a week. The group published more than 50 papers in two years and garnered unforeseen financial support.
Our relationship with the Institut Pasteur in France exceeded our expectations. QBI and the Institut Pasteur formed the Center for Emerging and Neglected Diseases in San Francisco in 2022.
QBI went from managing one lab to working with nearly two dozen across two continents. What role did technology play in facilitating this?
Fabius: Realizing technology's role in scientific communication was crucial. Globally, we formed a dozen, small specialized subgroups to streamline communications. Broadly, these looked at either the technology used to understand the virus or the biological processes hijacked during infection. We created similar groups on Slack and email.
It was important to make our communication technology facilitate the work, including empowering younger scientists, who played major roles during the pandemic.
The QBI Coronavirus Research Group released its findings initially as a preprint instead of in a peer-reviewed journal, why?
Fabius: We released our initial mapping online in March 2020 as a preprint, before it had been peer reviewed. To publish in academic journals, research has to be peer reviewed but it can take weeks if not months. Before the pandemic, there was skepticism about publicly sharing research prior to peer-review. During COVID-19, we joined others in publishing our work as preprints because it allowed information to be shared widely and quickly.
The formal peer review process remained an essential component of the scientific process. Still, in a fast-moving pandemic, the benefit of "crowd review" outweighed the risks.
What was the benefit of sharing information widely and quickly?
Krogan: When we released that paper on the preprint server bioRxiv, I tweeted out: "Hey, we have these plasmids, or copies of proteins that key for COVID-19 infection, that we made to study the virus."
"We're happy to send them to whoever, no strings attached." I added, "We'll even pay for shipping."
We sent our plasmids to about 400 labs in 42 countries to help expedite COVID-19 research. I like to say that these plasmids spread around the world much faster than the actual virus did.
Why did QBI partner with pharmaceutical companies, who usually focus on for-profit research and development?
Fabius: Pharmaceutical companies reached out to us. They weren't seeking transactional relationships, they were asking how they could contribute. The approach was unconventional and, for us, unprecedented. Partnering with industry drew initial skepticism but the academic community couldn't do this alone.
We're currently partnering with one company to test one of the drugs we mapped as a COVID-19 drug and to expand its uses in cancer care.
Rather than depend on scientific journals and media outlets, QBI took to YouTube, TikTiok and blogs to explain its work. Why?
Fabius: To succeed, scientists needed the public to understand their discoveries. To be heard amidst conspiracy theories and misinformation, we had to tell our own story so we learned the value of clear, consistent communication. Strategic use of blogs, Twitter, Instagram, Facebook and YouTube helped us to share information and make our science accessible.
Krogan: We regularly contacted media about discoveries and worked with public relations professionals around messaging. Our scientists also wrote personalized emails to donors. Effective communication and public exposure attracted supporters. We were particularly fortunate to find donors who supported us with unrestricted funds at a crucial juncture, which allowed the research to flourish.
What can we learn from COVID-19 to be better prepared for future pandemics?
Krogan: The problem with science, is that it's so siloed and it rewards the individuals over groups, for instance. What COVID-19 showed us is how fast we can move when we break down these silos across different labs, different institutions, disease focuses and even between academia and pharmaceutical companies.
Fabius: My question is, Why aren't we working on all diseases like this, including illnesses like breast cancer, Parkinson's or even HIV?
We live in a reality that undeniably will produce pandemics in coming years. Climate change is leading to shifts in temperatures, shrinking wild spaces and rises in people-made disasters that will put more people at risk of new diseases.
Krogan: My hope is that we can keep the collaborative research infrastructure we built during COVID-19 in place so we're much more ready for the next pandemic.
More information: Jacqueline M. Fabius and Nevan J. Krogan, Lessons learned for pandemic preparedness: A collaborative network is imperative, Cell Host & Microbe (2023). DOI: 10.1016/j.chom.2023.05.008. www.cell.com/cell-host-microbe … 1931-3128(23)00202-0