This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New sensor shows brain cells making and then breaking contact
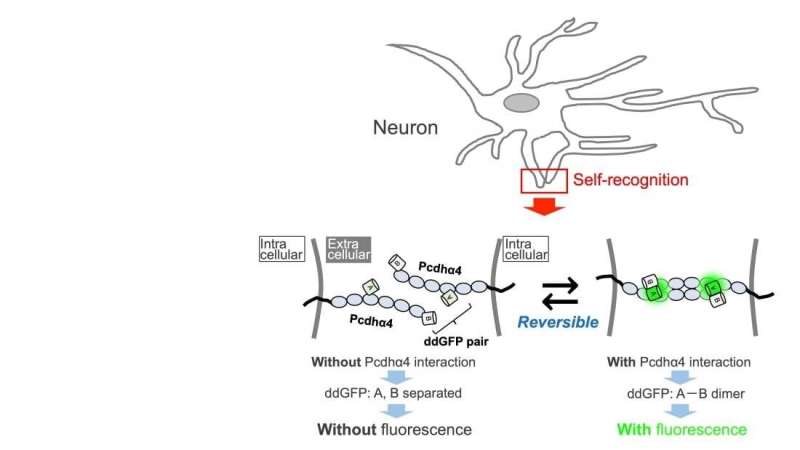
When brain cells, or neurons, are putting out processes to connect with other neurons, how do they tell the difference between their own processes and those of other neurons? One important part of this puzzle involves a molecule called clustered protocadherin (Pcdh).
In a recent publication in iScience, researchers from SANKEN (The Institute of Scientific and Industrial Research) and the Graduate School of Frontier Biosciences at Osaka University reported the development of a sensor to look at Pcdh interactions in live neurons, which brings us closer to understanding this mystery.
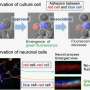
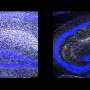
In the brain, millions of neurons make trillions of connections with one another. To do so, each neuron puts out tiny processes that grow and travel until they find another cell's processes to connect with. However, because each cell has so many processes all over the place, cells can accidentally make connections with themselves rather than with others. One way to avoid this involves Pcdh, which is expressed in different combinations on each neuron's surface.
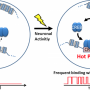
One role of Pcdh is in cell adhesion; if two neuronal processes have exactly the same combination of Pcdh molecules, the molecules bind to one another. Conversely, if the combinations are even slightly different, they are viewed as "other" rather than "self," and do not bind. There are conventional techniques for detecting molecular interactions between cell surfaces that can show us when the molecules bind, but not when they split apart again. Researchers from Osaka University wanted to tackle this issue.
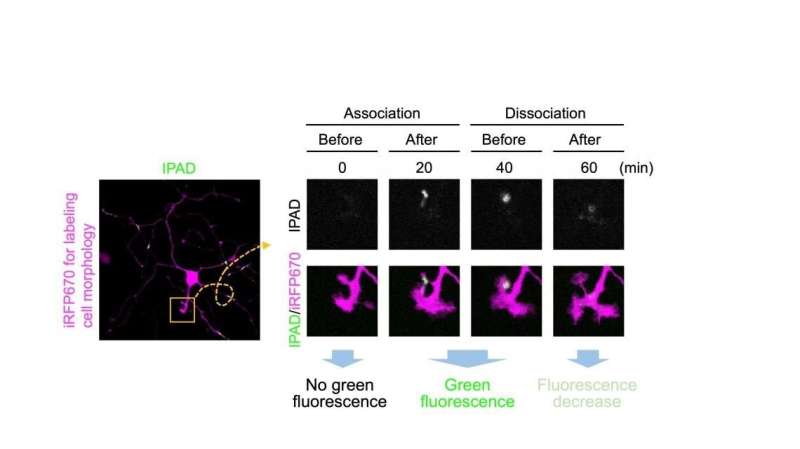
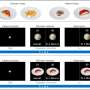
"We developed a fluorescent-based sensor that we named IPAD, or Indicators for Protocadherin Alpha 4 interactions upon Dimerization," says lead author of the study Takashi Kanadome. "This sensor allows us to see not only interactions between processes, but also the dissociation of these interactions for the first time."
This new technique does have a few disadvantages. For example, its fluorescence is much duller than that observed using older techniques, and it is unable to differentiate connections between processes from the same cell and those from two different cells with the same combinations of Pcdh on the surface.
"Despite its current drawbacks, we think that our new sensor will be useful for a number of different research applications," explains Tomoki Matsuda, senior author of the study. "The development of IPAD is an important step toward a better understanding of the neuronal recognition of self/other."
The sensors developed in this study have many potential applications. In particular, the technique may be used to develop a range of fluorescent sensors to visualize neuronal self-connectivity, which is implicated in brain disorders such as autism and epilepsy. A better understanding of neuronal self-connectivity may lead to improved treatments for these disorders.
More information: Takashi Kanadome et al, Visualization of trans-interactions of a protocadherin-α between processes originating from single neurons, iScience (2023). DOI: 10.1016/j.isci.2023.107238