This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Algorithmic blood test analysis will ease diagnosis of cancer types, guide treatment
Thanks to machine learning algorithms, short pieces of DNA floating in the bloodstream of cancer patients can help doctors diagnose specific types of cancer and choose the most effective treatment for a patient.
The new analysis technique, created by University of Wisconsin–Madison researchers and published recently in Annals of Oncology, is compatible with "liquid biopsy" testing equipment already approved in the United States and in use in cancer clinics. This could speed the new method's path to helping patients.
Liquid biopsies rely on simple blood draws instead of taking a piece of cancerous tissue from a tumor with a needle.
"Liquid biopsies are much less invasive than a tissue biopsy—which may even be impossible to do in some cases, depending on where a patient's tumor is," says Marina Sharifi, a professor of medicine and oncologist in UW–Madison's School of Medicine and Public Health. "It's much easier to do them multiple times over the course of a patient's disease to monitor the status of cancer and its response to treatment."
Cancerous tumors shed genetic material, called cell-free DNA, into the bloodstream as they grow. But not all parts of a cancer cell's DNA are likely to tumble away. Cells store some of their DNA by coiling it up in protective balls called histones. They unwrap sections to access parts of the genetic code as needed.
Kyle Helzer, a UW–Madison bioinformatics scientist, says that parts of the DNA containing the genes that cancer cells use often are uncoiled more frequently and thus are more likely to fragment.
"We're exploiting that larger distribution of those regions among cell-free DNA to identify cancer types," adds Helzer, who is also a co-lead author of the study along with Sharifi and scientist Jamie Sperger.
The research team, led by UW–Madison senior authors Shuang (George) Zhao, professor of human oncology, and Joshua Lang, professor of medicine, used DNA fragments found in blood samples from a past study of nearly 200 patients (some with, some without cancer), and new samples collected from more than 300 patients treated for breast, lung, prostate or bladder cancers at UW–Madison and other research hospitals in the Big Ten Cancer Research Consortium.
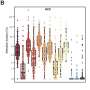
The scientists divided each group of samples into two. One portion was used to train a machine-learning algorithm to identify patterns among the fragments of cell-free DNA, relatively unique fingerprints specific to different types of cancers. They used the other portion to test the trained algorithm. The algorithm topped 80% accuracy translating the results of a liquid biopsy into both a cancer diagnosis and the specific types of cancer afflicting a patient.
In addition, the machine learning approach was able to tell apart two subtypes of prostate cancer: the most common version, adenocarcinoma, and a swift-progressing variant called neuroendocrine prostate cancer (NEPC) that is resistant to standard treatment approaches. Because NEPC is often difficult to distinguish from adenocarcinoma, but requires aggressive action, it puts oncologists like Lang and Sharifi in a bind.
"Currently, the only way to diagnose NEPC is via a needle biopsy of a tumor site, and it can be difficult to get a conclusive answer from this approach, even if we have a high clinical suspicion for NEPC," Sharifi says.
"Liquid biopsies have advantages," Sperger adds, "in that you don't have to know which tumor site to biopsy at, and it is much easier for the patient to get a standard blood draw."
The blood samples were processed using cell-free DNA sequencing technology marketed by Iowa-based Integrated DNA Technologies. Using standard panels like those currently in the clinic is a departure—one that can reduce the time and cost of testing—from other methods of "fragmentomic" analysis of cancer DNA in blood samples.
"Most commercial panels have been developed around the most important cancer genes that indicate certain drugs for treatment, and they sequence those select genes," says Zhao. "What we've shown is that we can use those same panels and same targeted genes to look at the fragmentomics of the cell-free DNA in a blood sample and identify the type of cancer a patient has."
The UW Carbone Cancer Center's Circulating Biomarker Core and Biospecimen Disease-Oriented Team contributed to the collection of the study's hundreds of patient samples.
More information: K.T. Helzer et al, Fragmentomic analysis of circulating tumor DNA-targeted cancer panels, Annals of Oncology (2023). DOI: 10.1016/j.annonc.2023.06.001